 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr2g008840.1 | ||||||||
| Common Name | MTR_2g008840 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1041aa MW: 116766 Da PI: 5.8218 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.5 | 4e-56 | 22 | 135 | 4 | 118 |
CG-1 4 ekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
++rwl++ ei++iL n++ +++t e++trp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v++l+cyYah+een++fqrr
Medtr2g008840.1 22 AQHRWLRPAEICEILRNYRMFHITPEPHTRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDALHCYYAHGEENENFQRR 115
49******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylevk 118
+ywlLe++ ++iv+vhylevk
Medtr2g008840.1 116 SYWLLEQD-THIVFVHYLEVK 135
********.*********985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 79.933 | 15 | 140 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.6E-76 | 18 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 6.7E-51 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 8.9E-8 | 453 | 529 | IPR002909 | IPT domain |
| CDD | cd00102 | 0.00458 | 453 | 541 | No hit | No description |
| SuperFamily | SSF81296 | 1.22E-17 | 454 | 539 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 4.3E-6 | 454 | 533 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF48403 | 2.95E-19 | 630 | 748 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.2E-8 | 636 | 715 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 20.022 | 637 | 758 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 2.6E-19 | 642 | 749 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 8.29E-16 | 643 | 740 | No hit | No description |
| SMART | SM00248 | 740 | 654 | 683 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.19 | 654 | 686 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.002 | 687 | 716 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.018 | 687 | 719 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 2300 | 726 | 755 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.41E-7 | 864 | 915 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.092 | 864 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.023 | 866 | 894 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0025 | 867 | 885 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0038 | 887 | 909 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.413 | 888 | 912 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 890 | 909 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1041 aa Download sequence Send to blast |
MAEPPSFGLG PRLDIQQLQF EAQHRWLRPA EICEILRNYR MFHITPEPHT RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDALH CYYAHGEENE NFQRRSYWLL 120 EQDTHIVFVH YLEVKSNKSN IGGNADSNEV ISDSQKVNSP SSGIPATYSS VPSLSTDSMS 180 PTSSYTSLRE DADSGDHGQS SVSGMDYIPP FSRDTFRGNG ATCIDGQASW DTVLQSTAEL 240 HADPSLVSFT SIPSGSLSNI LDQEDNILGD FSMSRSGLAI GAGSSQPLQS NWQIPFEDNT 300 GHMPTFTQSL SLEFASDYGT GLLGNESDNG SSIIDPVLFS FHGEPKEKLA QQNYLEEKVD 360 GHPRDDLKSN STKEVPSEET INYPLPVRRT LLDRDESLRK VDSFNRWITK ALGEVDDLNM 420 QSSPGISWSA DDCGHVIDDT SLSPSLSQDQ LYSITDFSPK WAYAESDTEV LIIGSFLKSQ 480 PDVTACNWSC MFGEVEVPAE VVANGILCCQ APPHKVGRVP FYVTCANRLA CSEVREFDFR 540 DGYSRNVDYT DFFNSSNDML LHLRLEEFLS LKPVHPSNQT FEGDTEKRSL ILKLISLREE 600 EEYSSKEEQT VEMDISRHKV KKHLFHRQFK EKLYSWLLHK VTESGKGPNV LDKDGQGVLH 660 LAAGLGYDWA IILILAAGVN INFRDVNGWT ALHWAASCGR ERTVGALVHM GADCGALTDP 720 SPEFPSGRTA ADLASSNGNK GLSGFLAESS LTSHLESLTV DDLHKGGQQE VSRTKAVQTV 780 SERTATPVIY NDMPDALCLK DSLTAVRNAT QAADRIHQVF RMQSFQRKQL TQDEDDDDEF 840 GLLDQRALSL LASKARKSGQ GDGLVNAAAT QIQKKFRGWK KRKEFLLIRQ RIVKIQAHVR 900 GHQVRKQYKT VIWSVGILEK IILRWRRKGS GLRGFRPEAL NKAPSQQNDS LKEDDYDYLK 960 EGRKQKEEKI QKALSRVKSM VQYPEARAQY RRVLNVVEDF RQKKDCNMGM SSEETVDGVE 1020 DLIDIDMLLD DENFNPIAFD * |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Mtr.8266 | 0.0 | leaf| root| stem | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, old leaves, petals, sepals, top of carpels, stigmas, stamen filaments, anthers and siliques, but not in pollen. {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
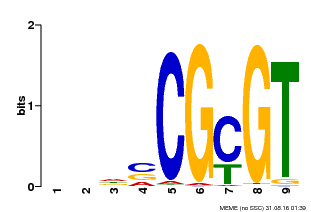
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr2g008840.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC187465 | 0.0 | AC187465.3 Medicago truncatula chromosome 2 BAC clone mth2-69g14, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003593198.2 | 0.0 | calmodulin-binding transcription activator 2 isoform X2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | G7INW0 | 0.0 | G7INW0_MEDTR; Calmodulin-binding transcription activator 1 | ||||
| STRING | AES63449 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.1 | 0.0 | ethylene induced calmodulin binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr2g008840.1 |
| Entrez Gene | 11433865 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



