 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g098900.1 | ||||||||
| Common Name | MTR_1g098900 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 638aa MW: 70838.1 Da PI: 5.823 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 93.3 | 2.4e-29 | 57 | 141 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW++qe+laL+++r++m++ +r+++ k+plW+evs+k++e g++rs+k+Ckek+en+ k++k++k+g+++++ + +t+++fdqlea
Medtr1g098900.1 57 RWPRQETLALLRIRSDMDTVFRDASVKGPLWDEVSRKLAELGYHRSSKKCKEKFENVYKYHKRTKDGRGGKS--DGKTYRFFDQLEA 141
8********************************************************************964..5557*******85 PP
| |||||||
| 2 | trihelix | 106 | 2.6e-33 | 454 | 539 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev+aLi++r++m+++++++ k+plWee+s +m+ g++r++k+Ckekwen+nk++kk+ke++kkr +e+s+tcpyf+ql+a
Medtr1g098900.1 454 RWPKVEVQALINLRTSMDNKYQENGPKGPLWEEISLAMKNLGYNRNAKRCKEKWENINKYFKKVKESNKKR-PEDSKTCPYFHQLDA 539
8*********************************************************************8.99***********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.018 | 54 | 116 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS50090 | 6.922 | 56 | 114 | IPR017877 | Myb-like domain |
| CDD | cd12203 | 1.88E-23 | 56 | 121 | No hit | No description |
| Pfam | PF13837 | 9.6E-19 | 56 | 142 | No hit | No description |
| PROSITE profile | PS50090 | 6.946 | 447 | 511 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0022 | 451 | 513 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-4 | 452 | 510 | IPR009057 | Homeodomain-like |
| CDD | cd12203 | 4.78E-29 | 453 | 518 | No hit | No description |
| Pfam | PF13837 | 1.6E-22 | 453 | 540 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 638 aa Download sequence Send to blast |
MLGDSAVLSG GGGDAPAPAA APPNMTVEGG GGVGSNSESD VERGRVEDGE RSFGGNRWPR 60 QETLALLRIR SDMDTVFRDA SVKGPLWDEV SRKLAELGYH RSSKKCKEKF ENVYKYHKRT 120 KDGRGGKSDG KTYRFFDQLE ALDHFHTNPS PQNISKPPQI SAPTPSQVVT TATVSLPITE 180 STMSQVVNTT LSIPHATVPS ISMPQNNIAT TQPIMNMNPT IPSNPSTIFQ PSTTNPTSTN 240 PLPSFPNIST DLLSNSMASS YSTSSEDTTE EGSRKRKRKW KNFFERIMKK VTEKQEDLQK 300 RFLEVIEKRE QERVVREEAW RAQEMQRINR EREMLAHERS ITAAKDAAVM SFLQKIAEQQ 360 NLGQALHNIN IAQPPPPQQQ LPQRSVAPTP TPAVVPISVV QVNTTPPPAQ PPPVSKLGTT 420 IVQQQQQQQL VTNMEIVKVD NNGETFMGGM SSSRWPKVEV QALINLRTSM DNKYQENGPK 480 GPLWEEISLA MKNLGYNRNA KRCKEKWENI NKYFKKVKES NKKRPEDSKT CPYFHQLDAL 540 YKEKGKGENY SGGGGGGAGS TQVVQPENMA APLMVQPEQQ WRPPQQGEDN MEQNRGQEEE 600 DMDEDDKDGE EEDDEEDMDE SGNFEVVANK PAVGASA* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 273 | 278 | RKRKRK |
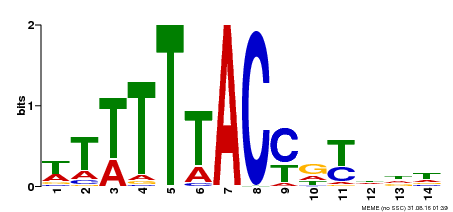
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g098900.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC225546 | 0.0 | AC225546.4 Medicago truncatula clone mth2-78i17, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003592114.1 | 0.0 | trihelix transcription factor GT-2 | ||||
| TrEMBL | G7ICZ1 | 0.0 | G7ICZ1_MEDTR; Putative transcription factor MYB family | ||||
| STRING | AES62365 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76880.1 | 3e-66 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g098900.1 |
| Entrez Gene | 11407016 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



