 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Medtr1g021520.1 | ||||||||
| Common Name | MTR_1g021520 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Medicago
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 297aa MW: 33316.4 Da PI: 9.1262 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.2 | 6.5e-32 | 127 | 180 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++Fv+av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
Medtr1g021520.1 127 PRMRWTTTLHSHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 180
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.63E-15 | 124 | 180 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-28 | 125 | 180 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-23 | 127 | 180 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.9E-7 | 128 | 179 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 297 aa Download sequence Send to blast |
MHTLFSPLLE PDLSLNISLP SNISDSEPKG ITNICSISTT SDSASSGSEL SHENPFIYPH 60 QRDPTLRLGF GNSDLMNPHH HNHHLHRHQV QGVSRNFNQH FQPHIYGRDF KRNTRVVNGV 120 KRSVRAPRMR WTTTLHSHFV HAVQLLGGHE RATPKSVLEL MNVKDLTLAH VKSHLQMYRT 180 VKTTDKSGAG HGILQTQGIN IVPLHGANSS ADERPNLPQP LQNSLRTSWQ PSIETNTNNT 240 EEKSEIGLTY SQLKENDTMV KRLDSAQLID LEFTLGRPNW GKDHAESSRE LSLLKC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-17 | 128 | 182 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: During ovule development, first expressed in the inner integument primordium and then on the abaxial side of the inner integument. {ECO:0000269|PubMed:16623911}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the periphery of the primary root apex and lateral root. {ECO:0000269|PubMed:15286295}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that regulates carpel integuments formation. Required for the specification of polarity in the ovule inner integument. Modulates the content of flavonols and proanthocyanidin in seeds. {ECO:0000269|PubMed:16623911, ECO:0000269|PubMed:19054366, ECO:0000269|PubMed:20444210}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
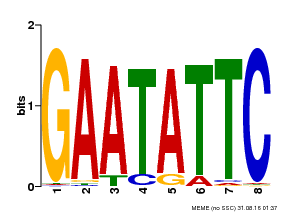
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Medtr1g021520.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC157978 | 0.0 | AC157978.25 Medicago truncatula clone mth2-47l16, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003589289.1 | 0.0 | probable transcription factor KAN4 | ||||
| Swissprot | Q9FJV5 | 5e-69 | KAN4_ARATH; Probable transcription factor KAN4 | ||||
| TrEMBL | G7I588 | 0.0 | G7I588_MEDTR; Myb-like DNA-binding domain, shaqkyf class protein | ||||
| STRING | AES59540 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4463 | 33 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.1 | 4e-67 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Medtr1g021520.1 |
| Entrez Gene | 11420402 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



