 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 69240 | ||||||||
| Common Name | MICPUCDRAFT_69240 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Mamiellaceae; Micromonas; Micromonas pusilla
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 234aa MW: 27201.3 Da PI: 12.6232 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 44.3 | 4.3e-14 | 18 | 62 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+ E+ +++a++++ + Wk+I ++g ++t q++s+ qky
69240 18 RESWTEKEHNMFLEAINMYDRD-WKKIETYVG-TKTVIQIRSHAQKY 62
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.92E-17 | 12 | 67 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.817 | 13 | 67 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 6.3E-18 | 16 | 65 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 8.0E-7 | 17 | 58 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.9E-11 | 17 | 65 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.1E-12 | 18 | 62 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.69E-9 | 20 | 63 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 234 aa Download sequence Send to blast |
MKPDASKARK PYTITKSRES WTEKEHNMFL EAINMYDRDW KKIETYVGTK TVIQIRSHAQ 60 KYFLKVRARQ IPPPRRTRRP SGRRRGRNVN RPPRRCSNHL MLDCFFFDTR HPPTSLSLSP 120 LPGPHRSKRT ARASTSRRRD RNASPRRRTR KNPPRAARHR AEGREPRLRR RRRKPPPPPP 180 RERERPRPPP RRPTRSAGTT STSTPRVGCK VRAGRGKGGA WARGATRSPA RGR* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 167 | 174 | LRRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
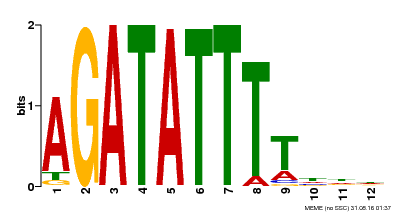
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003060779.1 | 1e-162 | predicted protein | ||||
| TrEMBL | C1MXY4 | 1e-160 | C1MXY4_MICPC; Predicted protein | ||||
| STRING | XP_003060779.1 | 1e-161 | (Micromonas pusilla) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP454 | 14 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.2 | 3e-31 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 69240 |
| Entrez Gene | 9685909 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



