 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010091969.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 192aa MW: 22042 Da PI: 5.7974 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 100.9 | 7.5e-32 | 131 | 189 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDg++WrKYG+K+vk+s++pr+YYrC+ +gCpvkk+ver++edp++v++tYeg Hnh+
XP_010091969.1 131 LDDGFKWRKYGKKMVKNSPNPRNYYRCSVEGCPVKKRVERDREDPRYVITTYEGVHNHQ 189
59********************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 3.9E-34 | 117 | 189 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.96E-28 | 124 | 189 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 30.813 | 126 | 191 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.8E-37 | 131 | 190 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 4.7E-25 | 132 | 189 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009867 | Biological Process | jasmonic acid mediated signaling pathway | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 192 aa Download sequence Send to blast |
MITNSKYYSY PYSSLINMSS GTSNNNNLVP MYQSPENDFA DTQSSFDPFP DLSVFDEWQL 60 DEDSEHPVLM PHDSGSMPHP TYHASEVHDE SGGHSSQFGG HAGRGENESG RERRDTKERV 120 AFKTQSEAEI LDDGFKWRKY GKKMVKNSPN PRNYYRCSVE GCPVKKRVER DREDPRYVIT 180 TYEGVHNHQS SF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 4e-25 | 121 | 188 | 7 | 74 | Probable WRKY transcription factor 4 |
| 2lex_A | 4e-25 | 121 | 188 | 7 | 74 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00531 | DAP | Transfer from AT5G26170 | Download |
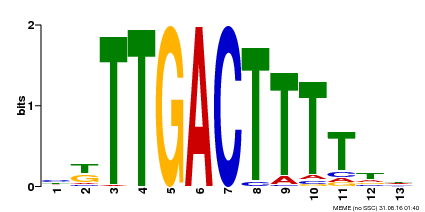
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010091969.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010091969.1 | 1e-143 | probable WRKY transcription factor 50 | ||||
| Swissprot | Q8VWQ5 | 8e-45 | WRK50_ARATH; Probable WRKY transcription factor 50 | ||||
| TrEMBL | W9R001 | 1e-142 | W9R001_9ROSA; Putative WRKY transcription factor 50 | ||||
| STRING | XP_010091969.1 | 1e-143 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1120 | 34 | 110 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G26170.1 | 2e-45 | WRKY DNA-binding protein 50 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21387306 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



