 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Migut.H00466.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Phrymaceae; Erythranthe
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 442aa MW: 49783 Da PI: 7.1041 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.7 | 9.7e-32 | 288 | 343 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+ F++a++qLGG+++AtPk+i+e+mkv+gLt+++vkSHLQkYRl+
Migut.H00466.1.p 288 KQRRCWSPELHRVFIDALQQLGGAQVATPKQIREVMKVEGLTNDEVKSHLQKYRLH 343
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.679 | 285 | 345 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-28 | 285 | 346 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.64E-17 | 286 | 346 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.2E-26 | 288 | 343 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.1E-6 | 290 | 341 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 442 aa Download sequence Send to blast |
MILHPHPLHA YLSGLCNNKT THTHTHTQRM GENYLKQVND SSCDGGSIFV AKTISDILTE 60 LSMIHDFSKK MSKLDFYIHS FQEEMRKIHV FQRELPLSMH LLKDAIERLK KESLKWEERE 120 RVPVMEEFIP LKSESSYDEG RAKPSNISQQ EAKKWMNSLH LWITPVQYEE DNNNQHSSFK 180 SKNKEEESSE SQSIFRNREG GAFSPFKKPS GVGVGVTIKE EKKAAAVVNP TVHGGGGGLT 240 LSIPVEADVA EHTTTAAQDS IVKDAPDQGI MIKKTKLKEP HQDEQQRKQR RCWSPELHRV 300 FIDALQQLGG AQVATPKQIR EVMKVEGLTN DEVKSHLQKY RLHVRKVPDP SSAPAARMNM 360 MGIIPWFRQD YEHGESSNRA KTNCFSPQQH LRDDMAGGST SAAQVGSATT TEEGESIEEA 420 DEDDQKSKSC SWKGKSAEHP F* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-13 | 288 | 341 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
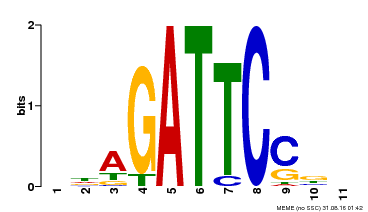
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Migut.H00466.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012856211.1 | 0.0 | PREDICTED: uncharacterized protein LOC105975554 | ||||
| TrEMBL | A0A022RWM5 | 1e-165 | A0A022RWM5_ERYGU; Uncharacterized protein (Fragment) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8497 | 22 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 2e-44 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Migut.H00466.1.p |



