 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.18G075600.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 223aa MW: 23871.8 Da PI: 5.2633 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 57.4 | 3.6e-18 | 27 | 77 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++ykGVr ++ +g+Wv+eIr p +++ r++lg++ t+e Aa+a++aa ++l+g
Manes.18G075600.1.p 27 KKYKGVRMRS-WGSWVSEIRAP---NQKVRIWLGSYSTPEAAARAYDAALLCLKG 77
59*****998.**********9...336************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.33E-20 | 27 | 87 | No hit | No description |
| PROSITE profile | PS51032 | 21.667 | 28 | 85 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.5E-11 | 28 | 77 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.0E-34 | 28 | 91 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.6E-29 | 28 | 85 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.24E-19 | 28 | 86 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 2.8E-8 | 29 | 40 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.8E-8 | 51 | 67 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0034605 | Biological Process | cellular response to heat | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071497 | Biological Process | cellular response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 223 aa Download sequence Send to blast |
MVKPENKIQS GTSKAMAATS PAAANKKKYK GVRMRSWGSW VSEIRAPNQK VRIWLGSYST 60 PEAAARAYDA ALLCLKGSSA NLNFPITSSH YIPDTVMSPK SIQKVAAAAA NSFMGNAAAA 120 ATASSPTPAT PSVSSPPLPS SFSSSPSPSS SISSTFSDQL DEDVSLVQSL GSYSETIPML 180 DDSWYNFEEL QSPKFLDQML NPPMTMMDDF YEGDIRLWSF AE* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-13 | 24 | 86 | 1 | 64 | ATERF1 |
| 3gcc_A | 1e-13 | 24 | 86 | 1 | 64 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in the regulation of plant development and tolerance to abiotic stresses (PubMed:21069430). Involved in salt and osmotic stress response pathways. May be regulated by the stress-related genes RD29A, RD22, DREB1A or P5CS during stress response (PubMed:27137403). Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250, ECO:0000269|PubMed:21069430, ECO:0000269|PubMed:27137403}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00156 | DAP | Transfer from AT1G21910 | Download |
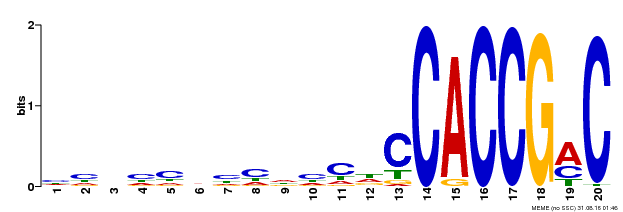
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.18G075600.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by jasmonate (JA) and ethylene. Down-regulated by freezing and heat stresses. {ECO:0000269|PubMed:21069430}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021600828.1 | 1e-160 | ethylene-responsive transcription factor ERF014-like | ||||
| Swissprot | Q9SFE4 | 5e-44 | ERF12_ARATH; Ethylene-responsive transcription factor ERF012 | ||||
| TrEMBL | A0A2C9U1F7 | 1e-159 | A0A2C9U1F7_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_026705m | 1e-160 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4798 | 31 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G21910.1 | 2e-31 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.18G075600.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



