 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.17G092200.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 567aa MW: 60992.8 Da PI: 9.7767 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.2 | 2.9e-05 | 66 | 88 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
Manes.17G092200.1.p 66 FICEVCNKGFQREQNLQLHRRGH 88
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 12.5 | 0.00043 | 142 | 164 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k + +s+ k H +t+
Manes.17G092200.1.p 142 WKCEKCSKKYAVQSDWKAHSKTC 164
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 5.38E-7 | 65 | 88 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.3E-6 | 65 | 88 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0052 | 66 | 88 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF12171 | 2.8E-5 | 66 | 88 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE profile | PS50157 | 10.99 | 66 | 88 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 68 | 88 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 150 | 107 | 137 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.1E-5 | 130 | 163 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.38E-7 | 137 | 162 | No hit | No description |
| SMART | SM00355 | 140 | 142 | 162 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 567 aa Download sequence Send to blast |
MAAASSSVSL FRSREENQNQ MMQQNSSTAA PTTAPQKKKR NQPGTPNPDA EVIALSPKTL 60 MATNRFICEV CNKGFQREQN LQLHRRGHNL PWKLKQKTTK EVKRKVYLCP EPTCVHHDPS 120 RALGDLTGIK KHYFRKHGEK KWKCEKCSKK YAVQSDWKAH SKTCGTREYR CDCGTLFSRR 180 DSFITHRAFC DALAQESARH PTSLSTIGTH LYGNNNMGIG LSQVGSQISS LQDQNHPTSN 240 MLRLGSTGAA KFEHIIPPSN SSSLSTMPSS AFFMSDANQG SFPNKSLHGL MQLPDLQSST 300 DNSSTTNLFN LGFFPSNATA PNRMNDSDNA NSSTVTTNLV NSRFLNSNPF HNGNGGQGTT 360 TLFANNMGDH VGSAGISSLY SNSMQQENIT PHMSATALLQ KASQMGSTTS SNNPNLLRSL 420 GSSPSTGIKS DRSPLVSTNF GSNSFGDATV GEVGLGIQME SDNQLQGLMN SLANGGSSIF 480 GGGHGQDNSF GGFTSGGVSL EQHHNSTKFR NVEEAKLHQS LGVGSDKLTL DFLGVGGMVR 540 SIGGGQHGIN LSSLDSQSAQ ANNKRF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 1e-34 | 138 | 199 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 1e-34 | 138 | 199 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
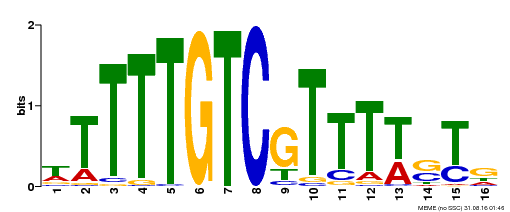
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.17G092200.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC208494 | 1e-144 | AC208494.1 Populus trichocarpa clone JGIACSB265-L05, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021599763.1 | 0.0 | protein indeterminate-domain 5, chloroplastic-like | ||||
| Swissprot | Q9ZUL3 | 1e-141 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A2C9U6M2 | 0.0 | A0A2C9U6M2_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_004443m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1536 | 33 | 97 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02070.1 | 1e-124 | indeterminate(ID)-domain 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.17G092200.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



