 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.15G149100.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 454aa MW: 51763.9 Da PI: 6.9831 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.9 | 4.2e-17 | 212 | 258 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT++Ed lv++vkq G + W+ Ia++m gR +kqc++rw+++l
Manes.15G149100.1.p 212 KGQWTPQEDRMLVHLVKQNGVKKWSQIAKMME-GRVGKQCRERWHNHL 258
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 59.7 | 6.2e-19 | 264 | 306 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++WT+eEde+l++a+k+ G++ W+ Ia++++ gRt++ +k++w+
Manes.15G149100.1.p 264 KDAWTEEEDEILIEAHKEIGNR-WAEIAKKLP-GRTENTIKNHWN 306
689*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 27.824 | 207 | 262 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.91E-32 | 210 | 305 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.4E-16 | 211 | 260 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.2E-16 | 212 | 258 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-27 | 213 | 265 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.02E-14 | 215 | 258 | No hit | No description |
| PROSITE profile | PS51294 | 22.171 | 263 | 313 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.3E-17 | 263 | 311 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 8.4E-17 | 264 | 306 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-23 | 266 | 311 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 7.62E-14 | 266 | 309 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 454 aa Download sequence Send to blast |
MELDRKLREE FPYLSSLLSD FPLKHEIESG FSSHEAPFSS NKGLFQSIHH LDHHNLHLPS 60 PFNSQYHLDH FTIEGSSKNP FLGVSATCID PLEPLPNGFS SDLNAFVSAA LLPANGGESG 120 YDHRPLHGSL QRRSFGDYNP QKFDEANDPL GQKLTYHHQS LNMRSMNLAK LPDEVSCITG 180 DNGYGKEADH RKDQRFQIKK DGKVHKKAQI IKGQWTPQED RMLVHLVKQN GVKKWSQIAK 240 MMEGRVGKQC RERWHNHLRP DIKKDAWTEE EDEILIEAHK EIGNRWAEIA KKLPGRTENT 300 IKNHWNATKR RQFTRRKGKE ANSKPTILQC YIKNLTSSSA TNHHQENNNP QDYLHKETSV 360 SSADHHHHLL KFPSSSLMHC DHNEGPNKFC VDTNFLFNDS YGFASSSLEE IPCTSVVDES 420 NLEYEISLEL YSLMKGAAAP AKEEMDLLEM ITQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 6e-45 | 210 | 312 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 6e-45 | 210 | 312 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
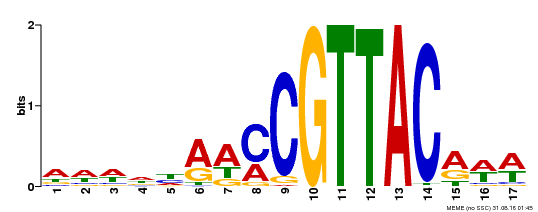
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.15G149100.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021595138.1 | 0.0 | transcription factor MYB98-like | ||||
| TrEMBL | A0A2C9UG32 | 0.0 | A0A2C9UG32_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_028573m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1570 | 31 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18770.1 | 2e-69 | myb domain protein 98 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.15G149100.1.p |



