 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.14G055900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1783aa MW: 201965 Da PI: 8.7732 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.6 | 0.0002 | 1666 | 1691 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sF +k +L++H r+ +
Manes.14G055900.1.p 1666 YQCDieGCSMSFGTKQELTVHKRNiC 1691
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 12.6 | 0.00041 | 1691 | 1713 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Manes.14G055900.1.p 1691 CPvkGCGKKFFSHKYLVQHRRVH 1713
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.4E-14 | 26 | 67 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.492 | 27 | 68 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 9.2E-15 | 28 | 61 | IPR003349 | JmjN domain |
| SMART | SM00558 | 5.2E-49 | 194 | 363 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.593 | 194 | 363 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.29E-26 | 212 | 381 | No hit | No description |
| Pfam | PF02373 | 2.3E-36 | 227 | 346 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 4.9E-5 | 1662 | 1688 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 6.8 | 1666 | 1688 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1689 | 1713 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.799 | 1689 | 1718 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.2E-6 | 1690 | 1717 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1691 | 1713 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.13E-9 | 1705 | 1747 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-8 | 1718 | 1743 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1719 | 1743 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1719 | 1748 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1721 | 1743 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.3E-8 | 1737 | 1771 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.6E-9 | 1744 | 1772 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.1 | 1749 | 1775 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.928 | 1749 | 1780 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1751 | 1775 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1783 aa Download sequence Send to blast |
MAASGGLVSE PSSQQEVFQW LKNLPLAPEY HPTLAEFQDP IAYIFKIEKE AAKYGICKIV 60 PPVLAAPRKA AIANLNRSLA ARAGSSSSKS PPTFTTRQQQ IGFCPRKPRP VQKPVWQSGE 120 NYTFQEFETK AKSFERSYLK KCSKKGAPSP LEIETLYWKA TVDKPFSVEY ANDMPGSAFS 180 PKKTGGKEVA EGVTVGETDW NMRGVSRAKG SLLRFMKEEI PGVTSPMVYV AMMFSWFAWH 240 VEDHDLHSLN YLHMGAGKTW YGVPREAAVA FEEVVRVHGY GGEMNPLVTF AILGEKTTVM 300 SPEVFVGAGV PCCRLVQNAG EFVVTFPRAY HSGFSHGFNC GEAANIATPE WLRVAKHAAI 360 RRASINYPPM VSHFQLLYDL ALELCTRMPL SISAKPRSSR LKDKQKGEGE TLVKELFIKN 420 VIHNNGLLHI LGKGSSIVLL PRSSSDISVC SNLRVGSQLR ASPALGSWSN KGIMKSSKDS 480 VPDEIMLERN NRINHAKGLF SVKEKFASLC GRNRFSSLDG NDNMNSTETG NENRESIHGD 540 KLSDQRLFSC VTCGILSFDC IAVIQPREAA ARYLMSADCS FFNDWMVGSG VTKDGFTIAH 600 GDTNTSEQNS STKWVEKNNV DGLYDVPVQS ANYQIQMMDQ NIVASNVETQ RATSALGLLA 660 LNYGNSSDSE DDQVEPDVSH QATEIDMTNC SSESKHQYQI SALPSFKQEF HHDTTGSHIV 720 SLSRHDNGHE VTLQTLDGHA EHGHGHMPAN FKDGSDQTLD SSVEFETDNL ASLESNGLEH 780 TFKDSMLTSL KTSSCSPVVH DTEKVVVPRE NTDESFAQRS DEDSSRMHVF CLEHAVEVEQ 840 QCRPIGGVHI LLLCHPEYPR IEAEAKLVTE ELGIDYFWND ITFRDATKED KDNIQSALDS 900 EEAIPGNGDW AVKLGINLFY SANLGRSSLY SKQMPYNSVI YKAFGRVSPA SLPTKFNVYR 960 RKPSKQKKVV AGRWCGKVWM SNQVHPFLTK QDSDDQDQEQ EQDRSFRGWT RPDEKLERKS 1020 ENIYKTETTL AARKSGRKRR MTVPSGPGKK VKCLDAEDAA SDESEEDVSH KQHTRVYSRK 1080 RTKRIEREVS LDSLEDDFHL HYEKRTHRNK QAKSVDREDA ISDDSLRCNT NQHRRTLRSK 1140 QVKYIESEND ISYAFADNKI NKQYSRIQRT RTNQAKYAQT VREISDDSLE GDIHEWHGRV 1200 PKSTLAKFTR EDAVSDDSPE ESSRRSVKRQ NRTSGGSQAK FIERDGEVSD DVLEENAYQQ 1260 HTGSRRSRES KFIDRESAVS DDQLEGNTYQ QRMRIFRTKQ AKISKTENAI SDDSSEENVQ 1320 QQRRGIPRRK RAKFVESEDA VSDDLLEDDT ALEHKRRTPR SRKAKVVGRE DAVSDDLLED 1380 NTHLEHRRMP RSRKAKVVGT EGVSDDLQDN RQWQPRKTPR GKQAQFIDSE DVSDDQLEED 1440 AHWQPRKLSR CKQAVSIERE DVSIDLEEGN THWQPKMTPS RKQAKFSESE DVSDDLQEDD 1500 NHWQPRKTPR GKQPKLIERE DAVLDDLVEE YSNKQQQRIL RSKQKKPVAL SKMKRGAIEL 1560 VKQGSSRPKK KENFRSIKQE KQMPETPRLR NGKAKHNARR VESRDEELEG GPSTRLRKRP 1620 SKASKESETK LKEKLQNSRK KVKNASSVKP LSGQKNVKNK VEEAEYQCDI EGCSMSFGTK 1680 QELTVHKRNI CPVKGCGKKF FSHKYLVQHR RVHLDERPLK CPWKGCKMTF KWAWARTEHI 1740 RVHTGARPYI CAEEGCGQTF RFVSDFSRHK RKTGHSVKKS RG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-78 | 13 | 384 | 1 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-78 | 13 | 384 | 1 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1629 | 1640 | KLKEKLQNSRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
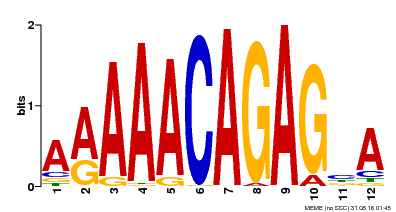
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.14G055900.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021634678.1 | 0.0 | lysine-specific demethylase REF6-like isoform X1 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2C9UK42 | 0.0 | A0A2C9UK42_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_000096m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.14G055900.1.p |



