 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.13G128300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 382aa MW: 42807.1 Da PI: 6.8886 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 105 | 4.4e-33 | 243 | 298 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++qLGGs++AtPk+i+elm+v+gLt+++vkSHLQkYRl+
Manes.13G128300.1.p 243 KQRRCWSPELHRRFVDALQQLGGSQVATPKQIRELMQVDGLTNDEVKSHLQKYRLH 298
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.367 | 240 | 300 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.99E-19 | 241 | 301 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-30 | 242 | 301 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.2E-27 | 243 | 298 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.6E-8 | 245 | 296 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010092 | Biological Process | specification of organ identity | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0048449 | Biological Process | floral organ formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 382 aa Download sequence Send to blast |
MELSLDLSLV YVPKTISEYL MEISGIKDSR RKLSKLDDYI KKLEDEMRKI DAFKRELPLC 60 MLLLNDAIVR LKQEALQCKE LEGEDRTGKQ EFVSVKENSG GDGGGNMGND LSDKKNWMSS 120 VQLWNTNNIN SDSKQHDSKS ETKQRSEEDD DQSTCENPVQ LCNYKSKGGA FMPFKALSVF 180 EGTERKEEKE VVSRVTDLSL MTPVSQLGSC NLISKCNANI QTKMHNKPQQ LQQHQQEHPS 240 YKKQRRCWSP ELHRRFVDAL QQLGGSQVAT PKQIRELMQV DGLTNDEVKS HLQKYRLHIR 300 KLPASSAACQ ASGLWMAQDH CKDPSKPSIS ESNSPQGPFH ACGYAKGISS TGGNSGEAED 360 DDKSESHSWT GRLHKAGEVD V* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 3e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 3e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 3e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 3e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 4e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
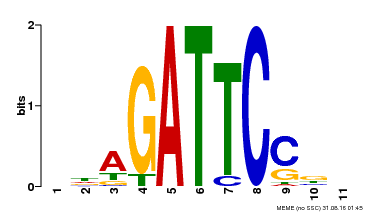
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.13G128300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021631588.1 | 0.0 | transcription factor HHO5-like isoform X1 | ||||
| TrEMBL | A0A2C9URI3 | 0.0 | A0A2C9URI3_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_009666m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1208 | 34 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37180.1 | 1e-78 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.13G128300.1.p |



