 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_AchrUn_randomP03000_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 601aa MW: 66708.8 Da PI: 6.7349 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.1 | 3e-16 | 72 | 116 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE+ ++++a k++G W +I +++g ++t+ q++s+ qk+
GSMUA_AchrUn_randomP03000_001 72 REKWTEEEHNRFLEALKLYGRS-WQRIEEHIG-TKTAVQIRSHAQKF 116
789*****************55.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.5E-16 | 66 | 122 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.838 | 67 | 121 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.4E-16 | 70 | 119 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.6E-13 | 71 | 119 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 7.0E-14 | 72 | 115 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.6E-9 | 72 | 112 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.30E-10 | 74 | 117 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 601 aa Download sequence Send to blast |
MRYFLDQVER LVGYMEGGWM IHVADFKSKC PRLLSTPPFS ASEVSDLEME EKSCGENMVA 60 KVRKPYTITK QREKWTEEEH NRFLEALKLY GRSWQRIEEH IGTKTAVQIR SHAQKFFTKL 120 EKEALIKGVS PGQVHDIDIP PPRPKRKPSN PYPRKMSTGV PSPSPSAEAT DDKPPTPSSL 180 QPTNQHSFRL ESDAATERLA AMETVAAKEN FEDGNRSEVI DINQDVPSAS FFTEFIPTSE 240 ASKKEKITVD KSLLLSEVNE EAVRGHLQHQ LPWFSPLTQC QSNQDACDSF LNISSTFSTL 300 IISTLLQNPS VHAAACLAAS VWPSPEVDTS LHSSPPVHMK PSPTMTPVVA ATVGAATAWW 360 AAQGALPWFP HFTFAFTSAP PTRAQPVEMV QEPERRHDAS FQVKASEINN SPPLLAGGVN 420 DLRSKRKQHL SCGSNDTANN ETVDKEAEEA FTNTSAPEIN HRRLRSSAST DEPWKESEKL 480 AYEAPLKRDL PRSSSPLHAK DAQTKVRSDR DSSMLPANLS ENAGLETDLL HPHCLKKEEM 540 SKEINGSTGQ SSVTNDFRHG NSMSSRTGFK PYKRCSVEAK ESKAVSGEET GNKKIRLPEE 600 A |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00103 | PBM | Transfer from AT2G46830 | Download |
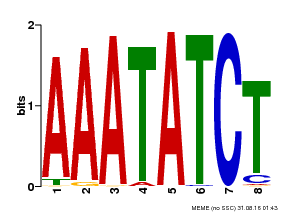
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009385738.1 | 0.0 | PREDICTED: protein LHY-like isoform X8 | ||||
| TrEMBL | M0U5W2 | 0.0 | M0U5W2_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_AchrUn_randomP03000_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01060.3 | 1e-40 | MYB_related family protein | ||||



