 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr4P01600_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 666aa MW: 72652.3 Da PI: 5.9992 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 100.4 | 1e-31 | 299 | 356 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK+vkgse+prsYY+Ct ++C+vkkkvers+ +++++ei+Y+g Hnh+k
GSMUA_Achr4P01600_001 299 EDGYNWRKYGQKQVKGSEYPRSYYKCTNPKCQVKKKVERSH-EGHITEIIYKGAHNHPK 356
8****************************************.***************85 PP
| |||||||
| 2 | WRKY | 102.7 | 2.1e-32 | 458 | 516 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDgy+WrKYGqK+vkg+++prsYY+Ct +gC+v+k++er+++d k v++tYeg+Hnh+
GSMUA_Achr4P01600_001 458 LDDGYRWRKYGQKVVKGNPNPRSYYKCTNPGCTVRKHIERASHDLKSVITTYEGKHNHD 516
59********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF118290 | 1.2E-24 | 296 | 357 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 2.8E-28 | 296 | 357 | IPR003657 | WRKY domain |
| SMART | SM00774 | 7.3E-35 | 298 | 356 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 4.1E-24 | 299 | 355 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 23.158 | 299 | 357 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 1.7E-36 | 443 | 518 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 3.92E-28 | 450 | 518 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 37.895 | 453 | 518 | IPR003657 | WRKY domain |
| SMART | SM00774 | 4.0E-37 | 458 | 517 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.9E-24 | 459 | 516 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 666 aa Download sequence Send to blast |
MAKTDDNNAM IDDRPPPNPS SRAFLSNSIN EEFGSKSFSD FLTENGNVGF RWTCENQKMG 60 INTKIEEAEA GNDFSNDASV QPKPFDAPKS CSAVGLAERM AARKGFNVPK LDTARIPPAT 120 IVSSSEICSP YLTIPPGLSP TMLLESPVFL ANCLAQPSPT TGKFNFADID SNPMSLSLSA 180 VSTKSDNDLF EDIPEAFSFK PPLESHSHLS STEEKVMLWM PGQTSHCKSC QFVSILILES 240 TQSVLYYILS TSRILAQLTH DKGFSDSIVG TDYAPTVDTQ QDGEADLREL SAAVGTPAED 300 GYNWRKYGQK QVKGSEYPRS YYKCTNPKCQ VKKKVERSHE GHITEIIYKG AHNHPKPHTS 360 RPPVAAEQCD PSNSLQQNQE GTHLSPEAIG VSSTMSNDEE EDDRATHGSV SLGCDGEGDE 420 TESKRRKLEA SAIEMSAASR AVREPRVVVQ TTSEVDILDD GYRWRKYGQK VVKGNPNPRS 480 YYKCTNPGCT VRKHIERASH DLKSVITTYE GKHNHDVPVA RNSGQPSSAQ SNTTANAAPQ 540 HNGLLQRPEP TQDGFVRFDG HAALGTFGFP GREQLGQPSS FPFSMAQPGL ANLAIAGLGP 600 MAAAMKMPVV PPLHPYLSHL QLTEAGLMVP KLEPKEESMP DSELPVLNAA SIYHQMMSRL 660 PLGPQL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 6e-37 | 299 | 518 | 8 | 77 | Probable WRKY transcription factor 4 |
| 2lex_A | 6e-37 | 299 | 518 | 8 | 77 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
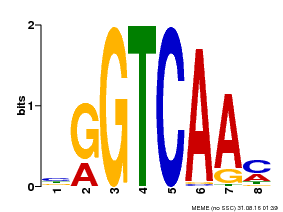
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | Transfer from AT5G56270 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009394983.1 | 0.0 | PREDICTED: probable WRKY transcription factor 2 isoform X2 | ||||
| TrEMBL | M0SK77 | 0.0 | M0SK77_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr4P01600_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2075 | 38 | 100 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56270.1 | 1e-154 | WRKY DNA-binding protein 2 | ||||



