 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | GSMUA_Achr10P03980_001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Zingiberales; Musaceae; Musa
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1497aa MW: 166667 Da PI: 9.0028 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.2 | 0.00027 | 1008 | 1031 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
y+C C++sFs+k +L H r
GSMUA_Achr10P03980_001 1008 YTCDieGCSMSFSTKQDLALHKRD 1031
99*******************985 PP
| |||||||
| 2 | zf-C2H2 | 12.4 | 0.00047 | 1033 | 1055 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H ++H
GSMUA_Achr10P03980_001 1033 CPvkGCGKKFFSHKYLVQHRKVH 1055
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11 | 0.0013 | 1091 | 1117 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
GSMUA_Achr10P03980_001 1091 YVCWesGCGQTFRFVSDFSRHKRKtgH 1117
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.7E-16 | 31 | 72 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.659 | 32 | 73 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.2E-15 | 33 | 66 | IPR003349 | JmjN domain |
| SMART | SM00558 | 2.1E-42 | 183 | 353 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 32.42 | 183 | 353 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.03E-22 | 197 | 347 | No hit | No description |
| Pfam | PF02373 | 1.8E-32 | 216 | 336 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 4.0E-4 | 1003 | 1030 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 7.3 | 1008 | 1030 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.49 | 1031 | 1055 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.967 | 1031 | 1060 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.52E-5 | 1032 | 1067 | No hit | No description |
| PROSITE pattern | PS00028 | 0 | 1033 | 1055 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-4 | 1033 | 1059 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| Gene3D | G3DSA:3.30.160.60 | 4.9E-8 | 1060 | 1084 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0026 | 1061 | 1085 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.72 | 1061 | 1090 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1063 | 1085 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.18E-9 | 1071 | 1115 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.4E-9 | 1085 | 1114 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1 | 1091 | 1117 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1091 | 1122 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1093 | 1117 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.710.10 | 1.6E-9 | 1171 | 1237 | No hit | No description |
| PROSITE profile | PS50097 | 9.889 | 1171 | 1232 | IPR000210 | BTB/POZ domain |
| SuperFamily | SSF54695 | 2.35E-9 | 1171 | 1237 | IPR011333 | SKP1/BTB/POZ domain |
| SMART | SM00225 | 6.1E-10 | 1171 | 1276 | IPR000210 | BTB/POZ domain |
| Pfam | PF00651 | 2.1E-7 | 1171 | 1272 | IPR000210 | BTB/POZ domain |
| Gene3D | G3DSA:3.30.710.10 | 1.6E-9 | 1276 | 1303 | No hit | No description |
| SuperFamily | SSF54695 | 2.35E-9 | 1276 | 1306 | IPR011333 | SKP1/BTB/POZ domain |
| CDD | cd14734 | 3.55E-14 | 1276 | 1335 | No hit | No description |
| SuperFamily | SSF57933 | 5.1E-15 | 1351 | 1459 | IPR000197 | Zinc finger, TAZ-type |
| Gene3D | G3DSA:1.20.1020.10 | 5.1E-17 | 1351 | 1460 | IPR000197 | Zinc finger, TAZ-type |
| SMART | SM00551 | 1.7E-8 | 1352 | 1451 | IPR000197 | Zinc finger, TAZ-type |
| Pfam | PF02135 | 5.0E-11 | 1359 | 1450 | IPR000197 | Zinc finger, TAZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0003712 | Molecular Function | transcription cofactor activity | ||||
| GO:0004402 | Molecular Function | histone acetyltransferase activity | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1497 aa Download sequence Send to blast |
MASAVSSSAQ GVVASEPPPM EVPPWLKSLP LAPEFHPTLQ EFQDPIAYIL KIEKEAADYG 60 ICKIVPPLPS APKKTAQQIG FCPRRPRPVQ KPVWQSGERY TLQQFEAKAR QFERSYLRRT 120 AGGGGRKAAT APALSPLEME TLFWRASADK PFSVEYANDM PGSGFAPLAA AAGRRCREEV 180 PANLGESAWN MRGVSRAKGS LLRFMKEEIP GVTSPMVYVA MLFSWFAWHV EDHELHSLNY 240 LHMGAGKTWY GVPRDAGLAF EEVVGVHGYG GDGNPLMTFA ILGEKTTVMS PEVFVGAGIP 300 CCSRLVQNAG DFVVTFPGSY HLGFSHGFNC GEAANIATPE WLKFAKGAAV RRASIDCPPL 360 VSHFQLLYAL ALSLCTRMPM CEGSEPRSSR LKDKIKGEGE EMVKNAFVQS VIQNNYLLSV 420 LLDSGNSCVV LPQNAPESPL CSTSLVRSQV KVKPRKSLGI CNFNGLFSFK GNFTSTGNDK 480 TVSLGTDNKF VNADHYSTSF ACVAVVQPRE TTAKYLMSAD CSFLDDHVSG SGEASGISRD 540 TNQRTNNNSL VADIVYCTAY PVQVSDQSVE MISDVTCPSG ASALDLLASI YEDSSDVEDE 600 DPENGKLVSA SYLEDNRTVA NSGTTTKFVG EPRGAQCREL DGQNAEDHYS NLKMGNLTSV 660 FKDLPVNLMQ GSDKDSSRLH VFCLEHAAEI EKQLQPIGGV HMMVLCHPDY PKIESEAKLL 720 AKEQGIGYIW KNVKFREANE EDQERIRAAI EDEQVMPMNS DWTVKLGINL CYTANLSKSP 780 LYSKQMPYNP VIYKAFGRDS TGNSPMKPKA TRRCPGRQKK IVAAGRWCGK VWMSNQVHPY 840 LAHKKETQEQ EQTEGLYSFD TDQNPLIEID IGHSSGLSSK RNSSGSNLAA NNSGKKRKRP 900 SKMAKSKKPL YSSTMADRNS KTDYVSAIPA SPLGRTLRSS HLRHNDSSSQ GKSSLKNESG 960 GPGTHLKKRS SKSEELKNKL ASKKQSTKRK AKNTQTASLA VKGKEEEYTC DIEGCSMSFS 1020 TKQDLALHKR DICPVKGCGK KFFSHKYLVQ HRKVHMDDRP LVCPWKGCKM TFKWPWARTE 1080 HIRVHTGDRP YVCWESGCGQ TFRFVSDFSR HKRKTGHSAK KGRRRSGLDR WVVEAQNIRR 1140 WLGNMTEEEK KRGEAEIQRW LEVGSVEIPP ADVRIVTSGG RRIPVHSTVL ASASPVLESI 1200 LGGPHKGRSR GREIPILGVP CDAVHAFVHS LYFARCVSAT EEEMIGEHGM HLLALSHAYR 1260 VGWLKRAAER ALSLRLTAEG VVDVLVLSQR CDAPRLHLRC MQLLSKDFAA VEQTEAWRFL 1320 QDHDPWLELH ILQFLHHAHL RRRRQARRKA EQRLYAELHE AMDCLRHICA DGCGEVGPSG 1380 RLLPADARPP CPNAVSCRGL RQLIRHFAAC DRQKRQQHGC RRCKRLWQLL RLHASICDEV 1440 DDASCKVPLC IQFKQRMQEK EGELEEDEDE RWRLLVKKVA SARVMSHLAK RQSQVLV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-71 | 25 | 442 | 8 | 413 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-71 | 25 | 442 | 8 | 413 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
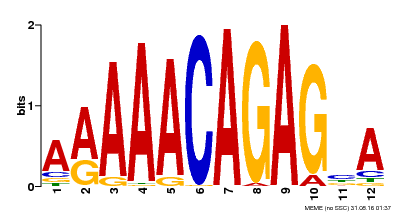
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020106930.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | M0RF75 | 0.0 | M0RF75_MUSAM; Uncharacterized protein | ||||
| STRING | GSMUA_Achr10P03980_001 | 0.0 | (Musa acuminata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5346 | 32 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



