 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LPERR01G03760.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Leersia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 276aa MW: 31055.8 Da PI: 6.1836 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58 | 2.4e-18 | 76 | 125 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
y+GVr++ +g+WvAeIr+p + r +r +lg+f ta eAa a+++a+++++g
LPERR01G03760.1 76 AYRGVRQRT-WGKWVAEIREP---N-RgRRLWLGSFPTAVEAAHAYDEAARAMYG 125
59****999.**********8...3.35************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF00847 | 1.2E-12 | 75 | 125 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 21.68 | 76 | 133 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 6.8E-34 | 76 | 139 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.05E-20 | 76 | 134 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 1.1E-30 | 77 | 134 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.5E-9 | 77 | 88 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 4.92E-31 | 77 | 135 | No hit | No description |
| PRINTS | PR00367 | 4.5E-9 | 99 | 115 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 276 aa Download sequence Send to blast |
MERGEVRTED CSPAQVRKKR TRRKSDGPDS IAETIKWWKE KNQKLQEENS SRKAPAKGSK 60 KGCMAGKGGP ENSHCAYRGV RQRTWGKWVA EIREPNRGRR LWLGSFPTAV EAAHAYDEAA 120 RAMYGSTARV NFADNSMDAN SGCTSAPSLM MSNGPTYVRS DEKDELESPP FMVTNCPPAV 180 LHQSAKKDVL ECVLPQVQDV KTEVSNDLRS IHEEQKTLEV CQSEGSVLHK EVSVTYDYFN 240 VHEVVEMIIV ELSADQKMEV HEEYKEGDDG FSLFSY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 2e-15 | 77 | 132 | 3 | 59 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]CCGAC-3'. Binding to the C-repeat/DRE element mediates high salinity- and dehydration-inducible transcription (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00302 | DAP | Transfer from AT2G40340 | Download |
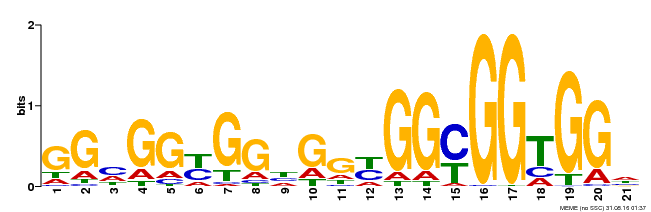
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LPERR01G03760.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF300971 | 0.0 | AF300971.2 Oryza sativa DRE binding protein 2 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006643786.1 | 1e-177 | PREDICTED: dehydration-responsive element-binding protein 2A | ||||
| Swissprot | A2WL19 | 1e-174 | DRE2A_ORYSI; Dehydration-responsive element-binding protein 2A | ||||
| TrEMBL | A0A0D9UX48 | 0.0 | A0A0D9UX48_9ORYZ; Uncharacterized protein | ||||
| STRING | LPERR01G03760.2 | 0.0 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7315 | 35 | 51 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40340.1 | 4e-35 | ERF family protein | ||||



