 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj2g3v1831990.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 356aa MW: 39009 Da PI: 8.4036 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.2 | 7.5e-33 | 220 | 275 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++qLGG+++AtPk+i++lm+v+gLt+++vkSHLQkYRl+
Lj2g3v1831990.1 220 KQRRCWSPELHRRFVDALQQLGGAQEATPKQIRDLMQVEGLTNDEVKSHLQKYRLH 275
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.851 | 217 | 277 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.27E-18 | 218 | 278 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-30 | 218 | 278 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.4E-26 | 220 | 275 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.2E-7 | 222 | 273 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010092 | Biological Process | specification of organ identity | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0048449 | Biological Process | floral organ formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 356 aa Download sequence Send to blast |
MDLCLDLSLA SSPSLFLGDV SRRSRDMSEK LAMLENYVQR LEDEMNKVEA FKRELPLCML 60 LVKEAIARLK EEKLQCLGKQ DPPAAAAAAA EESTPNCGAN GKGPGITMEK DKNNWMSTVQ 120 LWCAQTKSKN VEGDRSVPEK PIQLMNPNKT SVGGGFMTFI ADSRVPKEVS HVPSFKITPA 180 AQLSHSNSKS GCGGGSSSEL SQPQPQPPHH LLRQNQNPRK QRRCWSPELH RRFVDALQQL 240 GGAQEATPKQ IRDLMQVEGL TNDEVKSHLQ KYRLHVRRFP ISSNGQANNG LWMVQDQSGD 300 KSKGSLSQSG SPQGPLTPLL LGGSVKGLSS PGRNSADAED EQSDCWNWKG ADNQSL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 1e-14 | 220 | 273 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-14 | 220 | 273 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-14 | 220 | 273 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-14 | 220 | 273 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-14 | 220 | 273 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
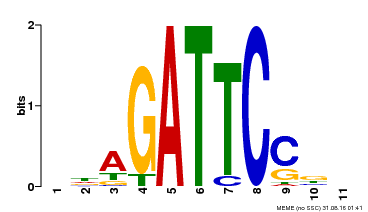
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj2g3v1831990.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT096930 | 1e-42 | BT096930.1 Soybean clone JCVI-FLGm-21I16 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027361911.1 | 1e-142 | transcription factor HHO5 | ||||
| TrEMBL | A0A371EC12 | 1e-134 | A0A371EC12_MUCPR; Transcription factor HHO5 (Fragment) | ||||
| STRING | XP_007156835.1 | 1e-128 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1208 | 34 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37180.1 | 3e-54 | G2-like family protein | ||||



