 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Lj1g3v1650270.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Loteae; Lotus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 414aa MW: 44529.7 Da PI: 7.9163 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 50.4 | 5e-16 | 351 | 404 | 5 | 58 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevak 58
+r+rr++kNRe+A rsR+RK+a++ eLe++v++L++eN++L+k+ ee+ ++ ++
Lj1g3v1650270.1 351 RRQRRMIKNRESAARSRARKQAYTMELEQEVAKLKEENEELQKKQEEIMEMQKN 404
69*********************************************9998765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.1E-13 | 347 | 411 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.645 | 349 | 400 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 4.0E-14 | 351 | 404 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 2.94E-11 | 351 | 401 | No hit | No description |
| CDD | cd14707 | 2.80E-27 | 351 | 405 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 6.2E-15 | 351 | 401 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 354 | 369 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 414 aa Download sequence Send to blast |
MNFKGFVNDP GGADNGNGGG RPPPGGFPLA RQSSVYSLTF DEFMTSMGGS GRDFGSMNMD 60 ELLKNIWSAE EVQSIGGGVG GGGEEGGGGA AVGHLQRQGS LTLPRTLSHK TVDQVWKDIS 120 KDYGPSLVAP PRQPTLGEMT LEEFLVRAGV VREDAKNDAV FADLARAGNN SGLGFEFQAQ 180 QMNRIAGLMG GNNRIPGASD DPIVSLQNST NLPLNANGFR SPNQQQQMQN SHQQNHQQQQ 240 HHQQQQIFPK QPALNYATQM PNQGIRGGIV GLSADQGMNG NLAQGGGIGG LVGLAPGPVQ 300 LASGSPANQM SSSDIMGKSN GDTSSVSPVP YVFNGGLRGR KTGAVEKVIE RRQRRMIKNR 360 ESAARSRARK QAYTMELEQE VAKLKEENEE LQKKQEEIME MQKNQVQEMM NLQR |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, leaves, flowers and siliques but not in seeds. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
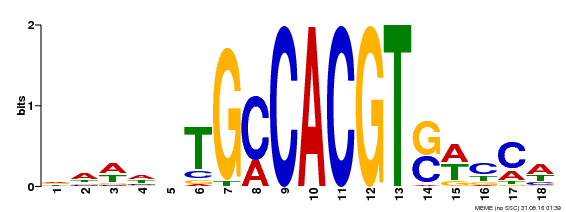
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Lj1g3v1650270.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003603049.1 | 0.0 | bZIP transcription factor 46 | ||||
| Refseq | XP_024635171.1 | 0.0 | bZIP transcription factor 46 | ||||
| Refseq | XP_024635172.1 | 0.0 | bZIP transcription factor 46 | ||||
| Swissprot | Q9M7Q4 | 1e-110 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | Q0PN11 | 0.0 | Q0PN11_9FABA; BZIP transcription factor | ||||
| STRING | AES73300 | 0.0 | (Medicago truncatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 7e-93 | abscisic acid responsive elements-binding factor 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



