 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0698s0007.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 337aa MW: 37207.5 Da PI: 4.4196 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 60.7 | 2.3e-19 | 96 | 148 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++++ +q++ Le+ Fe ++++ +++ +LAk+lgL+ rqV +WFqNrRa++k
Kalax.0698s0007.1.p 96 KRRLSVDQVQFLERSFEAENKLEPDRKIQLAKELGLQPRQVAIWFQNRRARWK 148
568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.9 | 5.1e-42 | 95 | 185 | 2 | 92 |
HD-ZIP_I/II 2 kkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
kkrrls +qv++LE+sFe+e+kLep+rK +la+eLglqprqva+WFqnrRAR+ktkq+Ekdy+aL+++y++lk++ ++L +e+e+L++e+
Kalax.0698s0007.1.p 95 KKRRLSVDQVQFLERSFEAENKLEPDRKIQLAKELGLQPRQVAIWFQNRRARWKTKQMEKDYDALQASYNSLKSSCDALVEENENLKAEV 184
9**************************************************************************************998 PP
HD-ZIP_I/II 92 k 92
Kalax.0698s0007.1.p 185 V 185
6 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.58E-19 | 82 | 152 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.054 | 90 | 150 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.4E-19 | 93 | 154 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.87E-19 | 95 | 151 | No hit | No description |
| Pfam | PF00046 | 1.4E-16 | 96 | 148 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-19 | 97 | 157 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 7.8E-6 | 121 | 130 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 125 | 148 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 7.8E-6 | 130 | 146 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 7.4E-17 | 150 | 191 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001558 | Biological Process | regulation of cell growth | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MAAGGSRVFR SGNLTSCSES MAPVSNPPAT ASSSRHSDAL FLPGSSRFPG SGSAMVSFED 60 VNGSMGRNTR SFFTAFDQDD NGDEDYEEYL NQAGKKRRLS VDQVQFLERS FEAENKLEPD 120 RKIQLAKELG LQPRQVAIWF QNRRARWKTK QMEKDYDALQ ASYNSLKSSC DALVEENENL 180 KAEVVALSEK LASKENPSQN AEPSEASKQS AAVPDNPISN SASEEEDSKF PLVEQKLLDD 240 FTSAKSDIFQ SDSPRYADGV YSPHLEHVDS SYVFEPDNNS DLSQDEDDSL PKAYLPSSSY 300 MLPPLKHGDY PDLSATSSSN FGSAHGDHQP FWSWSF* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 142 | 150 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00323 | DAP | Transfer from AT3G01470 | Download |
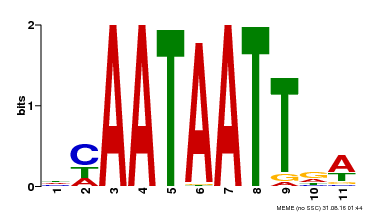
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022738207.1 | 1e-105 | homeobox-leucine zipper protein HAT5-like isoform X1 | ||||
| Refseq | XP_022738208.1 | 1e-105 | homeobox-leucine zipper protein HAT5-like isoform X2 | ||||
| TrEMBL | A0A067G8Q0 | 1e-104 | A0A067G8Q0_CITSI; Uncharacterized protein | ||||
| STRING | EOY21713 | 1e-104 | (Theobroma cacao) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G01470.1 | 3e-44 | homeobox 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0698s0007.1.p |



