 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kalax.0004s0169.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 249aa MW: 28114.2 Da PI: 7.0452 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58.9 | 1.1e-18 | 13 | 60 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg WT eEd++l ++v+ +G+g+W++Ia++ g++R++k+c++rw +yl
Kalax.0004s0169.2.p 13 RGLWTDEEDKILMEYVEVHGKGQWNRIAKKTGLKRCGKSCRLRWVNYL 60
788*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 61.1 | 2.3e-19 | 66 | 111 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+eE++l+++++k+lG++ W++Ia +++ gRt++q+k++w+++l
Kalax.0004s0169.2.p 66 RGNFTEEEEDLIIRLHKLLGNR-WSLIAGRVP-GRTDNQVKNYWNTHL 111
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.185 | 8 | 60 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.47E-31 | 11 | 107 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.2E-15 | 12 | 62 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.6E-17 | 13 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-25 | 14 | 67 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 9.06E-13 | 16 | 60 | No hit | No description |
| PROSITE profile | PS51294 | 26.858 | 61 | 115 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.7E-18 | 65 | 113 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.0E-18 | 66 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.86E-14 | 68 | 111 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-26 | 68 | 114 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010053 | Biological Process | root epidermal cell differentiation | ||||
| GO:0010091 | Biological Process | trichome branching | ||||
| GO:0048629 | Biological Process | trichome patterning | ||||
| GO:0090377 | Biological Process | seed trichome initiation | ||||
| GO:0090378 | Biological Process | seed trichome elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 249 aa Download sequence Send to blast |
MEGGGGVSHQ YRRGLWTDEE DKILMEYVEV HGKGQWNRIA KKTGLKRCGK SCRLRWVNYL 60 SPNVKRGNFT EEEEDLIIRL HKLLGNRWSL IAGRVPGRTD NQVKNYWNTH LSKRNGSTKR 120 PRTESYTPNP DTKSPACSKD SETMTAGAFV SLDPLRNNTG STTENPEQET TAAAALTTVN 180 NVLLDASFSL QNADAFLADG QPQHYASSSQ TWRDQMEQQM QMSSYQSLLQ FSDEFLQFDF 240 GWSSLFSD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 2e-28 | 13 | 114 | 7 | 107 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH2/EGL3/MYC146 or BHLH12/MYC1. Involved in epidermal cell fate specification in roots and hypocotyl. Together with GL3 or BHLH2, promotes the formation of non-hair developing cells (atrichoblasts) et the N position in root epidermis. Regulates stomata spatial distribution in hypocotyls. Binds to the WER-binding sites (WBS) promoter regions and activates the transcription of target genes such as GL2 and of CPC. {ECO:0000269|PubMed:10589676, ECO:0000269|PubMed:11585796, ECO:0000269|PubMed:14627722, ECO:0000269|PubMed:15361138, ECO:0000269|PubMed:15795220, ECO:0000269|PubMed:16207757}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00538 | DAP | Transfer from AT5G40330 | Download |
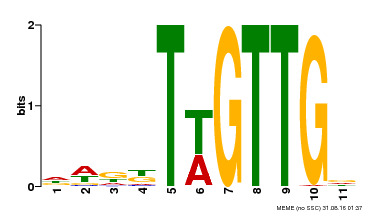
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Transcriptional activation correlates with reduced histone acetylation on H3 and H4 mediated by HDA18 in N cells. Repressed by CPC in hair cells (H position). {ECO:0000269|PubMed:11910008, ECO:0000269|PubMed:16176989}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010492291.1 | 6e-68 | PREDICTED: transcription factor WER-like isoform X1 | ||||
| Refseq | XP_015578183.2 | 1e-67 | transcription factor WER, partial | ||||
| Swissprot | Q9SEI0 | 2e-67 | WER_ARATH; Transcription factor WER | ||||
| STRING | Aquca_002_00170.1 | 3e-67 | (Aquilegia coerulea) | ||||
| STRING | XP_010492291.1 | 2e-67 | (Camelina sativa) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G14750.1 | 1e-68 | myb domain protein 66 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kalax.0004s0169.2.p |



