 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0089s0043.3.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 425aa MW: 46386.5 Da PI: 6.5072 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 32 | 2.2e-10 | 271 | 319 | 3 | 55 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L+el+P + +k Ka +L + ++Y++sLq
Kaladp0089s0043.3.p 271 NSHSLAERVRREKISERMKLLQELVPGC----NKITGKAVMLDEIINYVQSLQ 319
58**************************....999*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 4.97E-17 | 266 | 336 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.98E-11 | 266 | 323 | No hit | No description |
| PROSITE profile | PS50888 | 15.586 | 268 | 318 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 8.4E-17 | 269 | 335 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.5E-7 | 271 | 319 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-10 | 274 | 324 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 425 aa Download sequence Send to blast |
MGVGGDGEVD LGFHQNSNKG FMSCSTSGMN TNPVPNKAAG TAMSSIPMFK SSAGSDPFFG 60 THWDPLLAIN HSENFGGGSS MVSHCGYANP PYPIHNQGMS STHLVQYPSS DSTLVDLPKI 120 QHFGSGNFSD MAGLYGLPAL GATGTSSCYQ NYLPNQEVVT HRHHNTDGAR SQDDCQVSEE 180 RALEASPNGK RRRRSIESAS ELNSNKVTEG EQQKDMSEGS FDQPGEPDDK KQSQILKNKP 240 SSKQAKNNSP TEENTKDTYI HVRARRGQAT NSHSLAERVR REKISERMKL LQELVPGCNK 300 ITGKAVMLDE IINYVQSLQQ QVEFLSMKLA TVNPELDIDI ERMLSKDILQ SRGDNTVMLG 360 ISPGIRNSHP YPHAMLQSGM QNSATEYHHM PQNGWNNELQ SLLQMGFCSN PAVDNQGLDG 420 CSKP* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00135 | DAP | Transfer from AT1G10120 | Download |
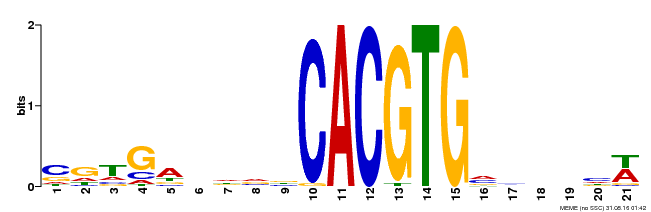
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021828015.1 | 1e-155 | transcription factor bHLH74 isoform X2 | ||||
| TrEMBL | A0A251RBG4 | 1e-152 | A0A251RBG4_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008220406.1 | 1e-152 | (Prunus mume) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G10120.1 | 4e-78 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0089s0043.3.p |



