 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Kaladp0063s0022.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Saxifragales; Crassulaceae; Kalanchoe
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 278aa MW: 30537 Da PI: 7.3011 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 29.6 | 1.6e-09 | 7 | 56 | 3 | 46 |
SS-HHHHHHHHHHHHHTTTT.......-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg.......tWktIartmgkgRtlkqcksrwqk 46
+W+ +Ed + da++ + + +W++Ia+ ++ +++ q+k++++
Kaladp0063s0022.1.p 7 SWSRDEDRAFEDAIATHWVEnledsdeHWEKIAAALP-TKSIHQLKHHYRL 56
7*****************99*****************.***********96 PP
| |||||||
| 2 | Myb_DNA-binding | 40.7 | 5.6e-13 | 126 | 170 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + +Rt+ q+ s+ qky
Kaladp0063s0022.1.p 126 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVITRTPTQVASHAQKY 170
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.499 | 3 | 62 | IPR017884 | SANT domain |
| SMART | SM00717 | 3.4E-7 | 4 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-5 | 6 | 57 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.0E-7 | 7 | 56 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.0E-7 | 7 | 64 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.37E-5 | 11 | 58 | No hit | No description |
| PROSITE profile | PS51294 | 18.403 | 119 | 175 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.38E-17 | 121 | 176 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.7E-17 | 123 | 173 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.3E-10 | 123 | 173 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-11 | 124 | 169 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.5E-11 | 126 | 170 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.65E-10 | 126 | 171 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 278 aa Download sequence Send to blast |
MTSSSSSWSR DEDRAFEDAI ATHWVENLED SDEHWEKIAA ALPTKSIHQL KHHYRLLVED 60 VNDIDAGIIP LPEYVGDDAS PSSSSKKSPY IADSNNNNNN SKKGFPGHES KANSNRSEQE 120 RRKGIPWTEE EHRLFLLGLD KFGKGDWRSI SRNFVITRTP TQVASHAQKY FIRLSSMNRD 180 RRRSSIHDIT SVNNGDITTP ITGQPASTNP NGGVPAGPTK HRPPEMGMYG APVGQVVGAP 240 VGHVAAVGTP VMMPPGHHPP YVVPVPYPVA PQPMHQH* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
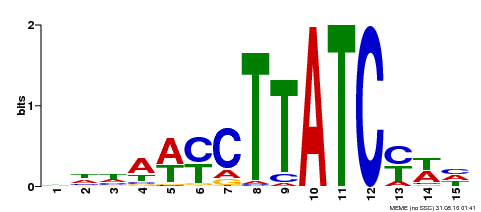
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021983810.1 | 1e-118 | transcription factor MYBS1 | ||||
| Swissprot | Q9FNN6 | 2e-82 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A251TYS3 | 1e-117 | A0A251TYS3_HELAN; Putative duplicated homeodomain-like superfamily protein | ||||
| STRING | VIT_17s0000g04130.t01 | 1e-111 | (Vitis vinifera) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G08520.1 | 7e-63 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Kaladp0063s0022.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



