 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00016051-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 560aa MW: 60950.8 Da PI: 6.5125 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 30.2 | 8e-10 | 364 | 411 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++Y++sLq
WALNUT_00016051-RA 364 SHSLAERVRREKISERMKLLQDLVPGC----NKITGKALMLDEIINYVQSLQ 411
8**************************....999*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 7.98E-17 | 358 | 430 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.22E-10 | 358 | 415 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 1.8E-16 | 360 | 428 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.656 | 360 | 410 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.5E-7 | 364 | 411 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 7.9E-10 | 366 | 416 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 560 aa Download sequence Send to blast |
MENEFFLNAG VPLSLHFESS ASMPMWHSLS SAMEIQGTEV NCLSEQSQDC FLNPNWEKSA 60 DHGVHFDSAL SSMVSSPAAS NTNLSNDSFV IRELIGKLGN IGNSGDISPH SQPLLASYIG 120 GNTSANTSCY STPLNSPPKL NVPRMDHLVK ENLPSLGKSM PLNSSVADFS ADPGFAERAA 180 RFSRFGSKSF NDRTSQFGIK NTDVSCRSYL PMDTEGLPRV SSSPSLNALG SQMGAQENNI 240 SPLQDRTELA NSQEESTVSE QIPKGETGLK GSNDVNSRKR KTASKGKAKE ATKAEEANDK 300 SNAKRCKTTE ANGNENGPVK AEEETKGSTR CAGDEKQSKS STKPPEPPKD YIHVRARRGQ 360 ATDSHSLAER VRREKISERM KLLQDLVPGC NKITGKALML DEIINYVQSL QRQVEFLSMK 420 LASVNTRMDF NMDSLMISKD IFRSNNPSPH AIFPLDSSAS AFYGHHPQQN PALHTNVSNG 480 TATQCPVDPL NTAPCQSLGV QLPPLNGYSE SVSQYPMYGE DDLQTIVQMG FGQNTNGETA 540 ILQPRRFHVS DQATHAKVEL |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 276 | 281 | SRKRKT |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00660 | PBM | Transfer from Pp3c6_9520 | Download |
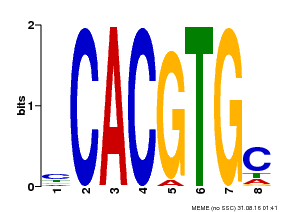
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018844170.1 | 0.0 | PREDICTED: transcription factor bHLH62 | ||||
| TrEMBL | A0A2I4GJU5 | 0.0 | A0A2I4GJU5_JUGRE; transcription factor bHLH62 | ||||
| STRING | EOX93799 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2530 | 34 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G07340.1 | 1e-118 | bHLH family protein | ||||



