 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | WALNUT_00014897-RA | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fagales; Juglandaceae; Juglans
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 642aa MW: 69679.7 Da PI: 4.5658 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 49.1 | 7.7e-16 | 219 | 255 | 1 | 35 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkgl 35
C++Cg ++ Tp++Rrgp g++tLCnaCGl+++ kg+
WALNUT_00014897-RA 219 CTHCGISSksTPMMRRGPAGPRTLCNACGLKWANKGV 255
*****99999************************997 PP
| |||||||
| 2 | GATA | 44.6 | 2e-14 | 497 | 531 | 1 | 33 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkk 33
C++Cg ++ Tp +Rrgp g+++LCnaCGl+++ k
WALNUT_00014897-RA 497 CQHCGISEksTPAMRRGPAGPRSLCNACGLMWANK 531
*****99999**********************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 14.448 | 78 | 113 | IPR010399 | Tify domain |
| SMART | SM00979 | 9.5E-12 | 78 | 113 | IPR010399 | Tify domain |
| Pfam | PF06200 | 5.4E-12 | 82 | 112 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.314 | 146 | 188 | IPR010402 | CCT domain |
| Pfam | PF06203 | 6.3E-16 | 146 | 188 | IPR010402 | CCT domain |
| PROSITE profile | PS50114 | 8.906 | 213 | 271 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 7.7E-12 | 213 | 266 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 4.75E-11 | 216 | 264 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.7E-14 | 218 | 259 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 3.27E-13 | 219 | 259 | No hit | No description |
| Pfam | PF00320 | 1.8E-13 | 219 | 255 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 219 | 246 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS51320 | 12.378 | 360 | 395 | IPR010399 | Tify domain |
| SMART | SM00979 | 9.6E-7 | 360 | 395 | IPR010399 | Tify domain |
| Pfam | PF06200 | 2.0E-8 | 363 | 394 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 12.759 | 428 | 470 | IPR010402 | CCT domain |
| Pfam | PF06203 | 1.3E-14 | 428 | 469 | IPR010402 | CCT domain |
| SMART | SM00401 | 4.6E-9 | 491 | 544 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 4.75E-9 | 492 | 535 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 5.9E-13 | 495 | 534 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 3.28E-11 | 496 | 547 | No hit | No description |
| Pfam | PF00320 | 2.1E-12 | 497 | 532 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 497 | 524 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 642 aa Download sequence Send to blast |
MPESNREASM YNSGAMNNNI ESHVEEEDED VAGDEESIDN PHMRFEDNAL AVKLNGGVQE 60 VATNSMYVPG SEFPPVAGNG GADQLTLSFQ GEVYVFDAVS PDKVQAVLLL LGGYEIPSVM 120 PTMGAAPLNL RVGMNDFPGR SIQPQRAASL SRFREKRKER CFDKKIRYSV RKEVALRMQR 180 KKGQFTSSKA IADVVGSESS VLSGAQGSGQ DDSMQETLCT HCGISSKSTP MMRRGPAGPR 240 TLCNACGLKW ANKGVLRDLP KVSNVSVLDS SVKTIEQGDG DAIDSDAVTP AAAELVSSAN 300 GDNSAPLQAR PFEEPEDQLV LVPIEVEGGS EDEYLDGGAP HEAMNVVEEV EARVNSGSVL 360 KSRTSELTVA FEGEVYVFPA VTPEKVQAVL LLLGGCDIPD SVPSSQFLLQ QNSRGIGDTS 420 RGSKLSRRIA SLVRFREKRK ERCFEKKIRY TCRKEVAQRM HRKNGQFASL KGTYKAAAEN 480 GDSSDGTPSP ATILRRCQHC GISEKSTPAM RRGPAGPRSL CNACGLMWAN KDTSADIKPS 540 TMEPENCYAD EDEQGSLNDA KPVPLEFKNP STRSGEQDFM QTTEVVTDHL SIELENPSVD 600 LGDQETIDEL ANASGTEFEI PENFDEQVDI DVSNMGTDWP RS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250, ECO:0000269|PubMed:14966217}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00369 | DAP | Transfer from AT3G21175 | Download |
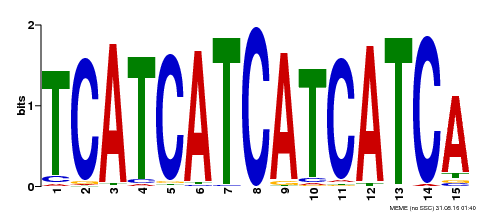
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018826887.1 | 0.0 | PREDICTED: GATA transcription factor 19-like | ||||
| Swissprot | Q8GXL7 | 2e-72 | GAT24_ARATH; GATA transcription factor 24 | ||||
| TrEMBL | A0A2I4F5G6 | 0.0 | A0A2I4F5G6_JUGRE; GATA transcription factor 19-like | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3319 | 33 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G21175.2 | 6e-74 | ZIM-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



