 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Itr_sc001303.1_g00003.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Convolvulaceae; Ipomoeeae; Ipomoea
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1338aa MW: 146920 Da PI: 8.9616 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.2 | 0.00026 | 1250 | 1272 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Itr_sc001303.1_g00003.1 1250 CPvkGCGKKFFSHKYLVQHRRVH 1272
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 12 | 0.00064 | 1308 | 1334 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Itr_sc001303.1_g00003.1 1308 YVCAepGCGQTFRFVSDFSRHKRKtgH 1334
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.1E-16 | 19 | 60 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.825 | 20 | 61 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 9.8E-15 | 21 | 54 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 29.684 | 180 | 393 | IPR003347 | JmjC domain |
| SMART | SM00558 | 2.4E-40 | 180 | 393 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.39E-22 | 195 | 260 | No hit | No description |
| Pfam | PF02373 | 1.8E-14 | 213 | 264 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.39E-22 | 307 | 411 | No hit | No description |
| Pfam | PF02373 | 8.9E-16 | 320 | 376 | IPR003347 | JmjC domain |
| SMART | SM00355 | 25 | 1225 | 1247 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 1248 | 1277 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1248 | 1272 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.0E-5 | 1249 | 1276 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1250 | 1272 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.62E-9 | 1264 | 1306 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.1E-8 | 1277 | 1302 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.741 | 1278 | 1307 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 1278 | 1302 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1280 | 1302 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.14E-8 | 1296 | 1330 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-9 | 1303 | 1331 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.385 | 1308 | 1338 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1308 | 1334 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1310 | 1334 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1338 aa Download sequence Send to blast |
MGEGSSGVEV LSWLKTLPVA PEYHPTLEEF QDPIAYIFKI EGEASKYGIC KIVPPVGMPP 60 KKTAIANLNK SLAARPHSTF TTRQQQIGFC PRRHRPVQKP VWQSGENYTL QQFEAKAKGF 120 EKNYLKKGSK KGLLPLEVES LYWKATVDKP FSVEYANDIP GSAFAPKKAD SVGAVGEGAT 180 VAETEWNMRG VSRARGSLLK FMKEEIPGVT SPMVYIAMMF SWFAWHVEDH DLHSLNYLHM 240 GAGKTWYGVP RDAAVAFEEV IRLQGYGGEI NPLGSIPLLC VHSDDAELGF PKVQIASLLL 300 AMTYFAASDL PPNVRLGVTF ATLGEKTTVM SPEVLLNAGV PCCRLVQDVG EFVVTFPRAY 360 HSGFSHGFNC GEASNIATPG WLRVAKDAAI RRASINCPPM VSHFQLLYDL ALSFCSSDPG 420 LCTPSQNAAC SIFNLLFEKM MSKNIRTEPR SSRLKDKKKD EGEALVKDFF VQDLEQNNSL 480 LHMLGRGSSV ILLPRNSTEN FITANFQGSQ FKVKPGLFSS VDCPDDAVET GKDLDLDYPL 540 IGRKQGMKQT LGVCSNKGKS SPLCESTRLP ISGRDSNVSS AIAHANGNMN TASERAYYNC 600 DRSSEHGLFS CVTCGILCYT CVAIVQPTKA ASRCLMSTDY ASFSQWREAG GGTTAVDADT 660 NVTMIGSSSG LMVRRPPDSL FDVSIKHTSQ LSDESVGVVS TSRTHKVSSS LGLLASTYGD 720 SSDSEADEAE ADVSVKDSEG RSMDCSPEDE IPLQLVIDPY ADHRQRRAQR DSETANRTSI 780 LSCSLQDRRA VPSCAPSAQK PPERAVKNAL VAPFGDAPMP SDEDSSRMHV FCLQHAVEVE 840 KELQLIGGAH ISLLCHPDYP KLEAQAKKIA KESGSDYPWN DILFQEATKD DEKLIRSALD 900 SEDAIHGNGD WAVKLGINLF YSANLSRSTL YRKQMPYNSV IYSAFGCSSP GGSPTSSENN 960 MKVPGKHKRI VVAGKWCGKV WMSNQVHSLL AERVSDEQEI RRNASSRFVP DGKPPPMRSL 1020 ECNQITETAT PTCRTGKKRK SVTESRPPAN SKSVKTDDDD PDETLSVGFS QLKSKTNLRN 1080 KRIKREAAAE PQKDSDKKNK GGGVDFVSED EVEGGPSTRL RRRAPKPAKQ QLETKPAKQQ 1140 LETKPIKAKP ALKKPEPKKP KQQENKKAKK GGGGPAMKVP AGNSSNSKKS PAVKNPPISN 1200 SVNKKGVLNK APSNSAKARD EDGEYPCDLE GCVMSFDSKQ ELVMHKKNIC PVKGCGKKFF 1260 SHKYLVQHRR VHMDDRPLKC PWKGCKMTFK WAWARTEHIR VHTGARPYVC AEPGCGQTFR 1320 FVSDFSRHKR KTGHSTKK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 8e-74 | 11 | 417 | 6 | 356 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 8e-74 | 11 | 417 | 6 | 356 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1154 | 1161 | PEPKKPKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
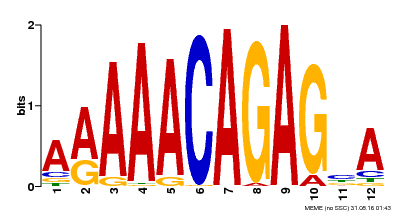
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019160805.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 isoform X2 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A484N386 | 0.0 | A0A484N386_9ASTE; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400039224 | 0.0 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4988 | 23 | 31 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



