 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_76271.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 363aa MW: 38665.4 Da PI: 8.3035 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 39.6 | 1.2e-12 | 107 | 151 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ ++ + ++lG+g+W+ I+r + Rt+ q+ s+ qky
MLOC_76271.1 107 PWTEEEHRMFLLGLQKLGKGDWRGISRSFVVSRTPTQVASHAQKY 151
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.60.10 | 2.3E-4 | 2 | 22 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS50158 | 8.565 | 3 | 19 | IPR001878 | Zinc finger, CCHC-type |
| PROSITE profile | PS51294 | 18.316 | 100 | 156 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.35E-16 | 102 | 157 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-18 | 103 | 155 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-10 | 104 | 154 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-11 | 104 | 150 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.07E-9 | 107 | 152 | No hit | No description |
| Pfam | PF00249 | 3.0E-10 | 107 | 151 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000122 | Biological Process | negative regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0030307 | Biological Process | positive regulation of cell growth | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:2000469 | Biological Process | negative regulation of peroxidase activity | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 363 aa Download sequence Send to blast |
MTRRCSHCSN NGHNARTCPA RSGGGVRLFG VHLTSPPVAA MKKSASMSCI ASSLGGGGSG 60 GSSPAAGPGP GGVARGGGEG APGYVSDDPM HASCSTNGRA ERKKGTPWTE EEHRMFLLGL 120 QKLGKGDWRG ISRSFVVSRT PTQVASHAQK YFIRQTNFSR RKRRSSLFDM VPEMPMDESP 180 DGAEEFTLCS TQDETTNSNK LSLFHLGRPK EAECDKDLPT LQLRQHEESE YAGRLLEAPD 240 FEMNNGVSFK AASVSTVPAF YPALLPVPLT LWPANVSNVE AANATHEVLK PTPVNVKEAI 300 KADEVVSMSK LSIGGDSSSS MEPSALSLQL TGPTNTRQSA FHVSPPMTRT DLSQGNNSPI 360 HAV |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00551 | DAP | Transfer from AT5G47390 | Download |
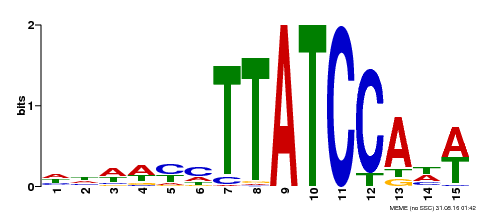
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_76271.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK371673 | 0.0 | AK371673.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv2139L02. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020189174.1 | 0.0 | transcription factor MYB1R1-like | ||||
| TrEMBL | F2E699 | 0.0 | F2E699_HORVV; Predicted protein | ||||
| STRING | MLOC_76271.3 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1939 | 37 | 91 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47390.1 | 4e-70 | MYB_related family protein | ||||



