 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MLOC_76165.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Hordeinae; Hordeum
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 685aa MW: 73998.1 Da PI: 6.8217 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 88.1 | 8.2e-28 | 215 | 268 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
+pr++W+ eLH++Fv av++L G++kA+Pk+ilelm+v+ Lt+e+v+SHLQkYR+
MLOC_76165.1 215 RPRVVWSVELHRKFVTAVNHL-GIDKAVPKRILELMNVEKLTRENVASHLQKYRV 268
59*******************.********************************7 PP
| |||||||
| 2 | Response_reg | 74.7 | 3.6e-25 | 26 | 134 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl vdD+p+ +++l+++l++ +y +v+++ ++ al++l+e++ +Dl++ D+ mp+mdG++ll+ e++lp+i+++ +ge +++ + + Ga
MLOC_76165.1 26 VLAVDDDPVCLKVLETLLRRCQY-HVTATNQAIIALRMLRENRdmFDLVISDVHMPDMDGFKLLELVGL-EMDLPVIMLSVNGETKTVMKGITHGAC 120
799********************.***************99999********************87754.558************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ +eel +
MLOC_76165.1 121 DYLLKPVRIEELKN 134
***********987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 2.2E-186 | 1 | 685 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 4.84E-34 | 23 | 148 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 1.1E-40 | 23 | 165 | No hit | No description |
| SMART | SM00448 | 3.2E-29 | 24 | 136 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.478 | 25 | 140 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 9.8E-23 | 26 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 5.83E-25 | 27 | 139 | No hit | No description |
| PROSITE profile | PS51294 | 11.6 | 212 | 271 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.5E-29 | 213 | 273 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 8.06E-19 | 213 | 272 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 7.1E-25 | 216 | 268 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.2E-6 | 217 | 267 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0031537 | Biological Process | regulation of anthocyanin metabolic process | ||||
| GO:0048367 | Biological Process | shoot system development | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0080036 | Biological Process | regulation of cytokinin-activated signaling pathway | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 685 aa Download sequence Send to blast |
MMRRDERRGA AAAATCRDQF PVGMRVLAVD DDPVCLKVLE TLLRRCQYHV TATNQAIIAL 60 RMLRENRDMF DLVISDVHMP DMDGFKLLEL VGLEMDLPVI MLSVNGETKT VMKGITHGAC 120 DYLLKPVRIE ELKNIWQHVV RRKFSTRDSL NADMYEECNK PPSSDSDYLY DQVKCGSSVQ 180 SGRPNKKRKE YHSEEEDEDG DSSGQDNDDP SAPKRPRVVW SVELHRKFVT AVNHLGIDKA 240 VPKRILELMN VEKLTRENVA SHLQKYRVYL RRLSAVASQQ AGIVAALGGR DPFLRMDAFE 300 GLQGYQAFTS PAALSTFGAH GLLNSPRNNQ TAVVIQGVAT SRSIPTGSSS CTTNPLIGDA 360 ANYHLSPPGN RHGNLAQGLV TSLGQTQLEQ KWMNEGSGDL SSILSGSSRA NGMPRMLPSV 420 TSSPLLPSDL VECTQAQVAI QPPVTAPSAS LEQLEGAIGA SSGTLVSRVS QQNALSPSGF 480 SANGLLVHGS LNNYGASFSS NGATRLGDAF SASFRQTNDF TVPRGMKVGS SSFGGAIHLS 540 PDTTEQKYLS FGSSDSLLDP KLVWSSSQLP SNIGAHHFMS QRSNNCSIGS SHGARMFGQT 600 SAGASTAVPQ AKFDILFSGD ASMARNASES SARRSESSAQ RLQSELSSST CSFDGLLNSM 660 IKVEKDDVAF SDDLGCDFYS FGACI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 2e-25 | 211 | 274 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00072 | SELEX | Transfer from AT4G31920 | Download |
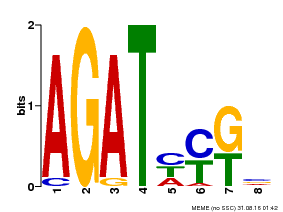
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | MLOC_76165.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK375849 | 0.0 | AK375849.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3107L06. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020170131.1 | 0.0 | two-component response regulator ORR23-like | ||||
| Swissprot | B8AEH1 | 0.0 | ORR23_ORYSI; Two-component response regulator ORR23 | ||||
| TrEMBL | M0Z328 | 0.0 | M0Z328_HORVV; Two-component response regulator | ||||
| STRING | MLOC_76165.1 | 0.0 | (Hordeum vulgare) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1856 | 38 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G25180.1 | 1e-105 | response regulator 12 | ||||



