 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | HL.SW.v1.0.G022183.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Humulus
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 507aa MW: 58361.7 Da PI: 5.7417 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 100.5 | 1.4e-31 | 51 | 135 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW++qe+laL+e+r++m+ ++r+++ k+plWeevs+kmre+gf+rs+k+Ckek+en+ k++k++++g++++++++ +++f+qlea
HL.SW.v1.0.G022183.1 51 RWPRQETLALLEIRSDMDAKFRDSSIKAPLWEEVSRKMREHGFNRSAKKCKEKFENIYKYHKRTRDGRSGKSNGKA--YRFFEQLEA 135
8********************************************************************9866665..*******85 PP
| |||||||
| 2 | trihelix | 104.8 | 6e-33 | 342 | 427 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k+ev aLi++r++++++++++ k+plWe++s++m++ g++rs+k+Ckekwen+nk++k++k+++k+r e+s+tcpyf+ql+a
HL.SW.v1.0.G022183.1 342 RWPKDEVDALIRLRTNLDTQYQDNGPKGPLWEDISAAMKKLGYNRSSKRCKEKWENINKYFKRVKDSNKRR-VEDSKTCPYFHQLDA 427
8********************************************************************98.78889********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.0044 | 48 | 110 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 1.7E-20 | 50 | 136 | No hit | No description |
| PROSITE profile | PS50090 | 7.039 | 50 | 108 | IPR017877 | Myb-like domain |
| CDD | cd12203 | 4.59E-25 | 50 | 115 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-5 | 338 | 398 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 9.1E-4 | 339 | 401 | IPR001005 | SANT/Myb domain |
| Pfam | PF13837 | 2.0E-22 | 341 | 428 | No hit | No description |
| CDD | cd12203 | 2.66E-24 | 341 | 406 | No hit | No description |
| PROSITE profile | PS50090 | 7.642 | 341 | 399 | IPR017877 | Myb-like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 507 aa Download sequence Send to blast |
MLENSTLPEN LSATTPENGE VGDSGGLAPV GHGEEERMGA EDGERSWLGN RWPRQETLAL 60 LEIRSDMDAK FRDSSIKAPL WEEVSRKMRE HGFNRSAKKC KEKFENIYKY HKRTRDGRSG 120 KSNGKAYRFF EQLEALDHHS FDHPFPPSGD KVNAFLDEAK TTAAATRTSP TDVVINAIPC 180 SIHKPASNLD DSSPSNTTSS SQESRGMRKQ KRRLTEFFER LMKEVTDSQE KLQRKFIETL 240 EKCEQDRLAR EEAWKKQELE RIKRESDLLV QERSIAAAKD AAVLAFLKKF SEQTEPFQFP 300 EYPIVDVDKQ EKNHSGNDEQ TCLDNQEKLN NNRKFVQVSS SRWPKDEVDA LIRLRTNLDT 360 QYQDNGPKGP LWEDISAAMK KLGYNRSSKR CKEKWENINK YFKRVKDSNK RRVEDSKTCP 420 YFHQLDALYN KKTKNVDDSE NTGYDLRPEE LLMHMMGGQE QQQQPETVTE NGESENLSHN 480 HEGDDEIVDK DEYKDTTDPS SMEIMG* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
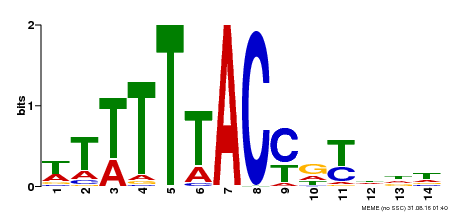
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010098893.1 | 0.0 | trihelix transcription factor GT-2 | ||||
| Swissprot | Q39117 | 1e-124 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A2P5ED76 | 0.0 | A0A2P5ED76_TREOI; TFIIB transcription factor | ||||
| STRING | XP_010098893.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76890.2 | 5e-66 | Trihelix family protein | ||||



