 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KDD72953.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Chlorellales
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 104aa MW: 11812.4 Da PI: 11.3931 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.2 | 8.9e-14 | 40 | 85 | 2 | 47 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
++W++ E+ +++++ + lG+g+Wk I+++ +Rt+ q+ s+ qk+
KDD72953.1 40 DPWSETEHTQFLHGLRALGKGQWKQISKYYVPTRTATQVASHAQKH 85
58***************************999************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.051 | 34 | 90 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.02E-16 | 35 | 90 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.9E-14 | 37 | 88 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.4E-9 | 38 | 88 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.4E-11 | 38 | 85 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.3E-11 | 41 | 85 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.27E-9 | 41 | 86 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000122 | Biological Process | negative regulation of transcription from RNA polymerase II promoter | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0030307 | Biological Process | positive regulation of cell growth | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:2000469 | Biological Process | negative regulation of peroxidase activity | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 104 aa Download sequence Send to blast |
MSGAPSPFQL PPRLALVDEM QRTKEQTEAT GSRRSTKKGD PWSETEHTQF LHGLRALGKG 60 QWKQISKYYV PTRTATQVAS HAQKHFLRVS GATKRRSRFT AMEQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to 5'-TATCCA-3' elements in gene promoters. Derepresses weakly the sugar-repressed transcription of promoters containing SRS. Contributes to the sugar-repressed transcription of promoters containing 5'-TATCCA-3' elements. {ECO:0000269|PubMed:12172034}. | |||||
| UniProt | Transcription activator that binds to 5'-TATCCA-3' elements in gene promoters. Derepress weakly the sugar-repressed transcription of promoters containing SRS. Contributes to the sugar-repressed transcription of promoters containing 5'-TATCCA-3' elements. {ECO:0000250|UniProtKB:Q7XC51}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00551 | DAP | Transfer from AT5G47390 | Download |
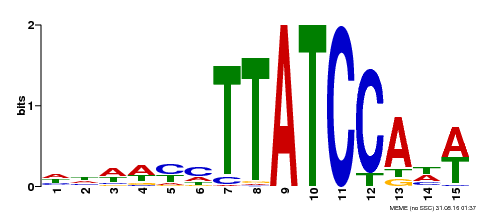
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by sucrose. Slightly repressed by gibberellic acid (GA). {ECO:0000269|PubMed:12172034}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011397494.1 | 2e-39 | Transcription factor MYB1R1 | ||||
| Swissprot | B8BI93 | 4e-20 | MYBS2_ORYSI; Transcription factor MYBS2 | ||||
| Swissprot | Q7XC51 | 4e-20 | MYBS2_ORYSJ; Transcription factor MYBS2 | ||||
| TrEMBL | A0A059LF47 | 2e-70 | A0A059LF47_9CHLO; Uncharacterized protein (Fragment) | ||||
| STRING | A0A087SFV2 | 9e-39 | (Auxenochlorella protothecoides) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP322 | 15 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47390.1 | 3e-22 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



