 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN38261.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 362aa MW: 42062.6 Da PI: 8.0756 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.7 | 9.9e-06 | 64 | 85 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg +F+ + +L +H+++H
KHN38261.1 64 TCEECGATFKKHAYLLQHMQSH 85
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 18 | 8.1e-06 | 91 | 115 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++C dC s++rk++L+rH+ H
KHN38261.1 91 FVCAvdDCQASYRRKDHLTRHLLQH 115
89*******************9887 PP
| |||||||
| 3 | zf-C2H2 | 14.5 | 0.0001 | 120 | 145 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kCp +C+ Fs +sn+krH H
KHN38261.1 120 FKCPveNCNSEFSLQSNMKRHTEEiH 145
89*******************87766 PP
| |||||||
| 4 | zf-C2H2 | 19.7 | 2.2e-06 | 161 | 185 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp Cgk+F+ s+L++H +H
KHN38261.1 161 HMCPeiGCGKVFRYASQLQKHEDSH 185
68*******************9888 PP
| |||||||
| 5 | zf-C2H2 | 16.7 | 2e-05 | 252 | 277 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C+ C+ +FsrksnL +H + H
KHN38261.1 252 FQCEfkGCSCTFSRKSNLDKHKKAvH 277
89*******************99877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50157 | 8.808 | 19 | 46 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 32 | 19 | 41 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.339 | 63 | 90 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.01 | 63 | 85 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.5E-8 | 64 | 90 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 65 | 85 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.17E-10 | 71 | 124 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.7E-10 | 91 | 117 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.018 | 91 | 115 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.39 | 91 | 118 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 93 | 115 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-5 | 118 | 146 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.12E-6 | 118 | 146 | No hit | No description |
| PROSITE profile | PS50157 | 10.201 | 120 | 150 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.029 | 120 | 145 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 122 | 145 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-7 | 156 | 185 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 6.25E-5 | 158 | 187 | No hit | No description |
| PROSITE profile | PS50157 | 10.907 | 161 | 190 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0014 | 161 | 185 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 163 | 185 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.7 | 193 | 218 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 195 | 218 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 30 | 221 | 242 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.8E-6 | 233 | 274 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.91E-9 | 234 | 291 | No hit | No description |
| PROSITE profile | PS50157 | 12.819 | 252 | 282 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0042 | 252 | 277 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 254 | 277 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 8.5E-5 | 276 | 306 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.3 | 283 | 309 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 285 | 309 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MCEAVEIEER PKYKDIRRYY CEYCGICRSK KTLITSHINL HHKEEYEKAR AERDPEAEAS 60 KSNTCEECGA TFKKHAYLLQ HMQSHSLERP FVCAVDDCQA SYRRKDHLTR HLLQHEGKIF 120 KCPVENCNSE FSLQSNMKRH TEEIHDDSST TTSSDKSHKQ HMCPEIGCGK VFRYASQLQK 180 HEDSHVNLES VDMVCLEPGC LKHFTNVQCL QAHVKSSHQY MTCDICGTKQ LKKNIKRHLR 240 SHEAEGSSSE TFQCEFKGCS CTFSRKSNLD KHKKAVHLNV KPFACGFPDC GMRFAYKHVR 300 DNHEKTAKHV FTLGDFEEAD EEFRSRPRGG RKRTCPTVEV LIRKRVTPPS QLEHWLFMQD 360 SE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 7e-15 | 67 | 220 | 19 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 7e-15 | 67 | 220 | 19 | 162 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
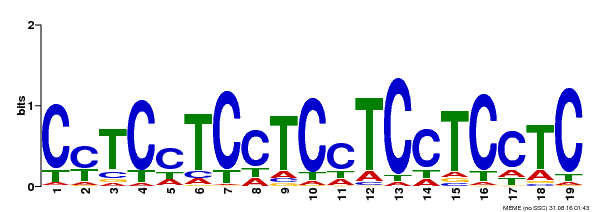
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN38261.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028220817.1 | 0.0 | transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 1e-140 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A445LVR3 | 0.0 | A0A445LVR3_GLYSO; Transcription factor IIIA isoform A | ||||
| STRING | GLYMA02G46270.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3069 | 33 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-109 | transcription factor IIIA | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



