 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KHN37514.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 439aa MW: 48341.9 Da PI: 10.119 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 44 | 4.8e-14 | 361 | 413 | 5 | 57 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkeva 57
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+ + ee+ ++ +
KHN37514.1 361 RRQRRMIKNRESAARSRARKQAYTFELEAEVAKLKELNRELQRKQEEIMEMKK 413
79************************************988877777777654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 5.6E-12 | 357 | 421 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.622 | 359 | 410 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.54E-9 | 361 | 409 | No hit | No description |
| Pfam | PF00170 | 1.2E-11 | 361 | 413 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 2.11E-26 | 361 | 415 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 1.7E-13 | 361 | 410 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 364 | 379 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 439 aa Download sequence Send to blast |
MNFRNFGVGD NHPTWDAMHG KTPANNNVTS TTLLRQPSTI YSLTFDEFQS TMGGIGKDFG 60 SMNMDELLKN IWTAEETQAM VFSAVAAAGG VEGHNNNSNN NPINCSGLQR QGSLTLPRTL 120 SQKTVEEVWR DLIKESGGEA NDGGSGGNGG SSNPQMQATL GEMTLEEFLV RAGVVREDVP 180 QQQQNGKPND NGWFGDFPRP NNNNTSLLLG FQQPNRSNGN GNLGENTNLV SKQQPPPLSL 240 NSNHSQRQAQ HQHQHQQHPP PLFPKPANVT FAGAPTHLLN NAHQHASPGR RGGLIGVAAE 300 HSMNVGMVGL ATANVTASPS SKISPDVITR SNNNVDNSPI SPHYVINRGR KFSAIEKVVE 360 RRQRRMIKNR ESAARSRARK QAYTFELEAE VAKLKELNRE LQRKQEEIME MKKNKDLDPA 420 CRPRISKIQC LRRTLTGPW |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Could participate in abscisic acid-regulated gene expression. | |||||
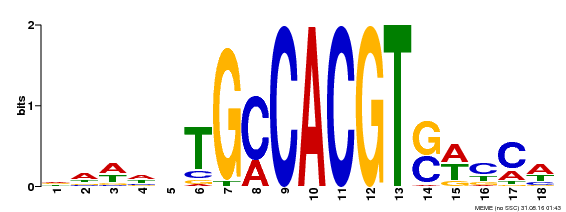
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KHN37514.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by abscisic acid (ABA) and cold. {ECO:0000269|PubMed:10636868}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT031133 | 0.0 | KT031133.1 Glycine max clone HN_CCL_105 BZIP transcription factor (Glyma02g14880.3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001238690.2 | 0.0 | bZIP transcription factor 1 | ||||
| Refseq | XP_025979754.1 | 0.0 | BZIP transcription factor bZIP1 isoform X1 | ||||
| Refseq | XP_028204869.1 | 0.0 | bZIP transcription factor TRAB1-like | ||||
| Swissprot | Q9M7Q5 | 6e-98 | AI5L4_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 4 | ||||
| TrEMBL | A0A0B2RZ62 | 0.0 | A0A0B2RZ62_GLYSO; ABSCISIC ACID-INSENSITIVE 5-like protein 4 | ||||
| TrEMBL | I1JET7 | 0.0 | I1JET7_SOYBN; BZIP transcription factor | ||||
| STRING | GLYMA02G14880.3 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49720.1 | 4e-80 | abscisic acid responsive element-binding factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



