 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.008G089900.4 | ||||||||
| Common Name | B456_008G089900, LOC105763524 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1047aa MW: 118044 Da PI: 6.5142 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 139 | 1.5e-43 | 14 | 95 | 37 | 118 |
CG-1 37 gsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
gsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een++fqrr+yw+Lee+l++ivlvhy++vk
Gorai.008G089900.4 14 GSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEENENFQRRSYWMLEEDLSHIVLVHYRDVK 95
9******************************************************************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01076 | 7.5E-48 | 1 | 95 | IPR005559 | CG-1 DNA-binding domain |
| PROSITE profile | PS51437 | 62.107 | 1 | 100 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.5E-36 | 14 | 94 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 5.1E-6 | 462 | 549 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.5E-6 | 463 | 548 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 8.4E-19 | 463 | 549 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 16.52 | 647 | 758 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.59E-12 | 647 | 756 | No hit | No description |
| SuperFamily | SSF48403 | 1.71E-14 | 648 | 759 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-14 | 648 | 759 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.217 | 697 | 729 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.03 | 697 | 726 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1000 | 736 | 765 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 8.23E-9 | 867 | 922 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.016 | 871 | 893 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.002 | 873 | 892 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.297 | 873 | 901 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0039 | 894 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 895 | 919 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.1E-4 | 897 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1047 aa Download sequence Send to blast |
MCCLSILLIL FLGGSLFLFD RKVLRYFRKD GHNWRKKKDG KTVKEAHERL KAGSIDVLHC 60 YYAHGEENEN FQRRSYWMLE EDLSHIVLVH YRDVKGNRTN FNRLKETEGG IPYSQEAVGI 120 VPNSEVESSL SSNFLPNNSQ IPSQTMDTAS MNSVHVQASE YEDAESVSNH QASSQFHSFL 180 DSHQPVAGRA DTRFYDPYVH VSHSNNCYGK PSITAFQLTQ TDKDREYNVA GITYEPQKNL 240 DFTSWEDVLE NCDRGVESAQ YQPPFTLKQN DTVGLLFDNS FLKKQAFEDQ SHAQEEWQGY 300 EGDSSHIVKW SLDQKLHPDL RYDLTSKFDE EVNHNLHPEK QHDHYLLNNQ LTDPSKGDHE 360 YVPKPDSENY LTLEGKSVYS SAMRPHLFDG SLAEGLKKLD SFNRWMSKEL GDVDESHTHS 420 SSGAYWDEVE GQNGIDVSSI PSQEQLDTFM LGPSLSHDQL FSIIDFSPNW AYVGSEIKVL 480 ITGRFLKSQG HAENCKWSCM FGEVEVPAEV IADGVLRCHT PKHEAGRVPF YVTCSNRLAC 540 SEVREFEYRV SHILDIDTVD NPSSNAIKIL DMRFGRLLCL GSSSPASNTN SIADISQLSS 600 KINSLLKEDV EEWDQMLAHN LEEDFFLEKL KEQLLKKLLK EKLRVWLLQK IVEGGKGPSI 660 LDKGGQGVIH FASALGYDWA LEPTIVAGVS VNFRDVNGWN ALHWAASSGR ERTVASLISL 720 GAAPGALTDP TPEYPLGRTP ADLASANGHK GISGYLAECD LSSHLLSLNL DKQGSASTTD 780 SRPDVIQKIL ELKTAPLNYG DASDGPSLKD SLAAVRNATQ AAARIHQVFR VQSFQNRQLK 840 EYGNDKYGMS DERALSLLAV KSNKPGQHDE RVHAAAIRIQ NKFRGWKGRK EFLIIRQRIV 900 KIQAHVRGHQ VRKNYRKIVW SVGIVEKVIL RWRRKGSGLR GFKPETLTKG PSVSVPSKED 960 DYDFLKKGRK QTEERLQKAL ARVKSMALNP AGRDQYSRIK NVVTEIQEKV LYDKVLNFAG 1020 ETTDLDKDLI DLEKLLDEDT FMQTAP* |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, carpels, and siliques, but not in stigmas or other parts of the flower. {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
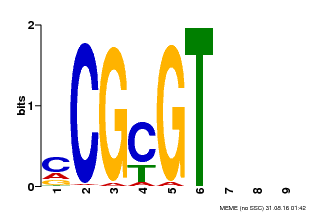
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AM434540 | 4e-39 | AM434540.2 Vitis vinifera contig VV78X210461.8, whole genome shotgun sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012437208.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X2 | ||||
| Refseq | XP_012437209.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X2 | ||||
| Refseq | XP_012437210.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X2 | ||||
| Refseq | XP_012437211.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A0D2TZT6 | 0.0 | A0A0D2TZT6_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.008G089900.1 | 0.0 | (Gossypium raimondii) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.008G089900.4 |
| Entrez Gene | 105763524 |



