 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.008G073300.1 | ||||||||
| Common Name | B456_008G073300, LOC105763342 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 367aa MW: 40939 Da PI: 6.7776 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 104.9 | 4.5e-33 | 230 | 285 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFvea++qLGGs++AtPk+i+elm+v+gLt+++vkSHLQkYRl+
Gorai.008G073300.1 230 KQRRCWSPELHRRFVEALQQLGGSQVATPKQIRELMQVDGLTNDEVKSHLQKYRLH 285
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.019 | 227 | 287 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.8E-30 | 227 | 288 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.38E-19 | 228 | 288 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.4E-27 | 230 | 285 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.5E-8 | 232 | 283 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010092 | Biological Process | specification of organ identity | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0048449 | Biological Process | floral organ formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 367 aa Download sequence Send to blast |
MELSLDLSLV YVPKTISEFL KEVSKIKNGF QRLAKITDYL QRLEDEMKKI DAFKRELPLC 60 MLLLKDGIER LKEEEIQCKE MNDGSVTEEN GRETMDNDGG DKKNWMSSVQ LWNSNFNNVD 120 PHNKPNTVPE LKLTSEEGED GSESENPIEI CNKQRKGGAF VPFKEQADKK VSGLSLMTPS 180 SELASCILKN NGGCRIGSGS SLYAQQNQIK FQTKSQIRHE EQPQQQNSRK QRRCWSPELH 240 RRFVEALQQL GGSQVATPKQ IRELMQVDGL TNDEVKSHLQ KYRLHIRKLP ASSSGQGNEL 300 CSAQNQCNVK MKASMSQSGS PQGPLLGSAS AKDMSSTGGD SMDTEDEKSD GHSWRSGIKK 360 PGEVDV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 8e-15 | 230 | 283 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 8e-15 | 230 | 283 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 8e-15 | 230 | 283 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 8e-15 | 230 | 283 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-14 | 230 | 283 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
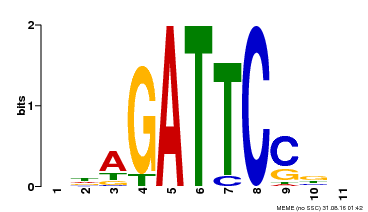
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX580318 | 1e-125 | JX580318.1 Gossypium hirsutum clone NBRI_GE12725 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012436995.1 | 0.0 | PREDICTED: uncharacterized protein LOC105763342 | ||||
| TrEMBL | A0A0D2PQG9 | 0.0 | A0A0D2PQG9_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.008G073300.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4922 | 26 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37180.1 | 2e-70 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.008G073300.1 |
| Entrez Gene | 105763342 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



