 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.003G041500.3 | ||||||||
| Common Name | B456_003G041500 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 301aa MW: 34094.4 Da PI: 4.5549 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.6 | 5.1e-19 | 45 | 98 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ +q+++Le+ Fe ++++ e++ +LA++lgL+ rqV vWFqNrRa++k
Gorai.003G041500.3 45 KKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWK 98
566899***********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.3 | 1.5e-41 | 44 | 136 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreel 91
ekkrrls +qvk+LE++Fe e+kLeperKv+la+eLglqprqvavWFqnrRAR+ktkqlE+dy Lk++y++lk++ e+L++++e+L +e+
Gorai.003G041500.3 44 EKKRRLSVDQVKALEKNFEVENKLEPERKVKLAQELGLQPRQVAVWFQNRRARWKTKQLERDYGLLKTSYETLKANYEALQHDNEALLKEI 134
69*************************************************************************************9999 PP
HD-ZIP_I/II 92 ke 93
+e
Gorai.003G041500.3 135 RE 136
86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.44E-19 | 30 | 102 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.346 | 40 | 100 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-17 | 43 | 104 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 6.71E-17 | 45 | 101 | No hit | No description |
| Pfam | PF00046 | 3.3E-16 | 45 | 98 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.7E-20 | 47 | 106 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.2E-6 | 71 | 80 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 75 | 98 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.2E-6 | 80 | 96 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 8.1E-16 | 100 | 140 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 301 aa Download sequence Send to blast |
MNDLCFGADE HSPRNNHVYS REFQSMLDGL DEEGCLEESG HVAEKKRRLS VDQVKALEKN 60 FEVENKLEPE RKVKLAQELG LQPRQVAVWF QNRRARWKTK QLERDYGLLK TSYETLKANY 120 EALQHDNEAL LKEIRELQAK HSGESTETES NVSVKEEVIV SENTDNKTLE QSEPAADLNY 180 GSVVLGATLF PDMKDGSSDS DSSAILNEDI ILNTNTNKDN SPNNAAFSSS GVLELDSQHH 240 LLISSSSSPT SSMDCFNFTK TSYQPHQYVK MEEHNFFTAD EACNFFSDPS LHWYSPEQWN 300 * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 92 | 100 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gra.2833 | 0.0 | flower|seedling| flowering | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
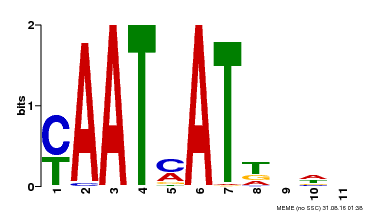
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ527086 | 0.0 | HQ527086.1 Gossypium herbaceum clone NBRI_D2_391 simple sequence repeat marker, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012469828.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-6-like | ||||
| TrEMBL | A0A0D2QN29 | 0.0 | A0A0D2QN29_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.003G041500.1 | 0.0 | (Gossypium raimondii) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 1e-59 | homeobox protein 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.003G041500.3 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



