 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gorai.001G239000.1 | ||||||||
| Common Name | B456_001G239000, LOC105766233 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 305aa MW: 34688 Da PI: 7.8396 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58 | 2.2e-18 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++lv +++q+G+g+W++++ + g+ R+ k+c++rw +yl
Gorai.001G239000.1 14 KGPWTPEEDLILVSYIQQHGPGNWRAVPTKTGLLRCSKSCRLRWANYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 47.7 | 3.6e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+ E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Gorai.001G239000.1 67 RGNFTENEEKMIIHLQALLGNR-WAAIASYLP-ERTDNDIKNYWNTHL 112
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-24 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 23.903 | 9 | 65 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.57E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.5E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.4E-16 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.07E-10 | 16 | 61 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.9E-24 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-14 | 66 | 114 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 19.48 | 66 | 116 | IPR017930 | Myb domain |
| Pfam | PF00249 | 2.6E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.40E-10 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001666 | Biological Process | response to hypoxia | ||||
| GO:0009617 | Biological Process | response to bacterium | ||||
| GO:0009626 | Biological Process | plant-type hypersensitive response | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042761 | Biological Process | very long-chain fatty acid biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 305 aa Download sequence Send to blast |
MGRQPCCDKV GVKKGPWTPE EDLILVSYIQ QHGPGNWRAV PTKTGLLRCS KSCRLRWANY 60 LRPGIRRGNF TENEEKMIIH LQALLGNRWA AIASYLPERT DNDIKNYWNT HLKKKLQGNE 120 SSSRYGFPSS SSSYQICRGQ WERKLQTDIH MAKKDLSDAL SPEKSSDLVE MKPFNNHTSS 180 PKPSGYASST ENIAKLLKGW MRNNPWKMAD SADYSEEGTV PMKEDKNSKE MAEAFQSHLG 240 FESLDSSLSD ISPSMSPETS LSQYESKPHL NAQSQLSLLE KWLFDEGKDY QLCDITLDQN 300 LYFF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gv2_A | 9e-26 | 14 | 116 | 4 | 105 | MYB PROTO-ONCOGENE PROTEIN |
| 1mse_C | 1e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| 1msf_C | 1e-25 | 14 | 116 | 4 | 105 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed in 2-week-old seedlings, in the early stages of development. {ECO:0000269|PubMed:10571865}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in vascular tissues of leaves, hypocotyl and roots. {ECO:0000269|PubMed:24587042}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to the DNA sequence 5'-AACAAAC-3' (PubMed:19170933). Acts as a positive regulator of hypersensitive cell death (PubMed:10571865, PubMed:12119395). Acts as a positive regulator of salicylic acid synthesis (PubMed:16730712). Regulates very-long-chain fatty acid biosynthesis (PubMed:18326828). Acts cooperatively with BZR2 to promote expression of a subset of brassinosteroids target genes (PubMed:19170933). Transcriptional activity and hypersensitive response control negatively regulated by PLA2-ALPHA and by the Xanthomonas type III effector XopD (AC G9L9K6) (PubMed:20696912, PubMed:21917550). Involved in the regulation of abscisic acid (ABA) signaling (PubMed:22814374). Increased levels of MYB30 can accelerate flowering both in long and short days through the regulation of FT (PubMed:24587042). {ECO:0000269|PubMed:10571865, ECO:0000269|PubMed:12119395, ECO:0000269|PubMed:16730712, ECO:0000269|PubMed:18326828, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20696912, ECO:0000269|PubMed:21917550, ECO:0000269|PubMed:22814374, ECO:0000269|PubMed:24587042}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00386 | DAP | Transfer from AT3G28910 | Download |
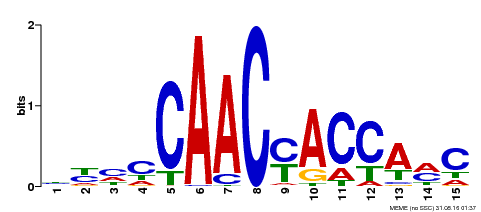
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated during hypersensitive response, but no expression detected during compatible interaction with pathogens (PubMed:10571865). Specifically induced in the inoculated zone 4 hours post pathogen infection (PubMed:20696912). Up-regulated by jasmonic acid and salicylic acid (PubMed:16463103, PubMed:16730712). Transcriptionally regulated by BZR2 (PubMed:19170933). {ECO:0000269|PubMed:10571865, ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:16730712, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20696912}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ118243 | 0.0 | DQ118243.1 Gossypium hirsutum myb transcription factor MYB30 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012441068.1 | 0.0 | PREDICTED: myb-related protein 306 | ||||
| Swissprot | Q9SCU7 | 1e-109 | MYB30_ARATH; Transcription factor MYB30 | ||||
| TrEMBL | A0A0D2PW31 | 0.0 | A0A0D2PW31_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.001G239000.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5842 | 23 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G28910.1 | 1e-108 | myb domain protein 30 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Gorai.001G239000.1 |
| Entrez Gene | 105766233 |



