 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KXZ48880.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Volvocaceae; Gonium
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 877aa MW: 89205 Da PI: 4.2812 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 54 | 2.3e-17 | 231 | 262 | 1 | 32 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrk 32
C+ Cg+t+Tp+WR+gp g+ktLCnaCG++y +
KXZ48880.1 231 CVECGATQTPQWREGPAGPKTLCNACGVRYGR 262
*****************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd04370 | 1.09E-9 | 34 | 159 | No hit | No description |
| SMART | SM00439 | 3.4E-4 | 35 | 170 | IPR001025 | Bromo adjacent homology (BAH) domain |
| PROSITE profile | PS51038 | 15.147 | 35 | 170 | IPR001025 | Bromo adjacent homology (BAH) domain |
| Pfam | PF01426 | 5.9E-5 | 36 | 164 | IPR001025 | Bromo adjacent homology (BAH) domain |
| PROSITE profile | PS50114 | 12.479 | 225 | 260 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.6E-12 | 225 | 279 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 6.1E-14 | 228 | 265 | IPR013088 | Zinc finger, NHR/GATA-type |
| SuperFamily | SSF57716 | 7.6E-12 | 228 | 266 | No hit | No description |
| CDD | cd00202 | 4.24E-12 | 230 | 260 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 231 | 256 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 8.8E-15 | 231 | 262 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 877 aa Download sequence Send to blast |
MSSAQVEFFF IGPPLPSTGP TTERLDFAGF SLNGIEYRVG DCAFLFPEDE SQGQHYVAQV 60 LSCFQLTRGQ AADPHCVEVA WFERRSHLDV ADVSGLDDEH NPQSLAKEVV ALEETDVNPI 120 GCISGKALVL KAPCYEEACR MATSCGMDSV DWFFCRGVYK QSEKCIIPFP EFADLLAANN 180 GKRLGKRAKP GLAGQVEVVA DSDDVKRQRF VEEQPAAPMR RTGKLVSGRT CVECGATQTP 240 QWREGPAGPK TLCNACGVRY GRAQQRANKR AVSSGLSRPS CGSGARNGRT SKTSRSEAAP 300 PPRASVRSAA AHAAEETSQR PLRQAALMSA SRTAAYARTG VFPLNSEELE MQLMEGDDGV 360 EVEGVERDAD PPLAVPQAAP LAGGPACRGS SGSGASPHNS SLDASSDSPA QTEPAEAMGP 420 AQAPLEVEVL VDGEADASVA AAAAAAGCPF APLPVLPYQL MPAHAPLLLS PSVFTDSGVV 480 PSAAVDAATA ISFGCETPLP LPPAPVAAAA VGFADAGLMS ALPCVPCDPS AGMIPEAEAD 540 ACAWMSMGLA MADDGAGPCG ADDQLRELQD AAAAADDEEP ALAGPAVSGD GRCASASCAD 600 ASCAHIVSAP VSPSARASCG GADCGLPAGM AGCGSCYGGG GSMSDDMCFY GGCSGVGSLD 660 DDEYTPLVAV EVDFGLGGCM AGSDLFDEVQ LQAGPAAECV PVSGAADLPA LAAAVERYSP 720 DTLAPGTIAR SVLDCLPALL APPEHQQPAA PATAPAQAPL LSEAAVAPLV VLGRQASRAS 780 CEAVAADAAY QAVSQVWEAH QEANRRAREV AAASLGRLNT ALSHLATGSG VDGVLVGAAG 840 APPPTAASHL IDLRDLCGGG MGAYGRDGAE MVVAVRS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
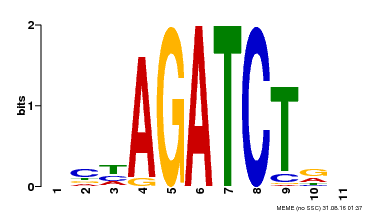
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A150GHQ1 | 0.0 | A0A150GHQ1_GONPE; Uncharacterized protein | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G32890.1 | 2e-14 | GATA transcription factor 9 | ||||



