 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.14G217700.1.p | ||||||||
| Common Name | GLYMA_14G217700, LOC100790289 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 931aa MW: 103499 Da PI: 5.8387 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75.5 | 6e-24 | 148 | 249 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f+k+lt sd++++g +++p++ ae+ +++++ ++l+++d++ ++W++++iyr++++r++lt+GW+ Fv +++L++gD+v+F +
Glyma.14G217700.1.p 148 FCKTLTASDTSTHGGFSVPRRAAEKLfppldyTIQPPT-QELVVRDLHDNTWTFRHIYRGQPKRHLLTTGWSLFVGSKRLRAGDSVLFIR 236
99*********************999*****9555555.49************************************************5 PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
d ++ +l+v+v+r
Glyma.14G217700.1.p 237 D--ERSQLRVGVRRV 249
5..55666***9985 PP
| |||||||
| 2 | Auxin_resp | 114.2 | 1.2e-37 | 274 | 357 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalk.vkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
aahaa+++s+F+++YnPra++seFv++++k++k++ ++vsvGmRf m+fete+s +rr++Gt+vg+sd+dp+rWp+SkWr+++
Glyma.14G217700.1.p 274 AAHAAANRSPFTIFYNPRACPSEFVIPLAKYRKSVFgTQVSVGMRFGMMFETEESGKRRYMGTIVGISDVDPLRWPGSKWRNIQ 357
79********************************988********************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.01E-46 | 137 | 277 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.3E-40 | 141 | 262 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 4.28E-23 | 147 | 248 | No hit | No description |
| PROSITE profile | PS50863 | 12.844 | 148 | 250 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.6E-21 | 148 | 249 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 1.5E-24 | 148 | 250 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.2E-32 | 274 | 357 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 23.555 | 824 | 908 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 8.34E-8 | 826 | 903 | No hit | No description |
| Pfam | PF02309 | 1.4E-7 | 870 | 911 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009908 | Biological Process | flower development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0010305 | Biological Process | leaf vascular tissue pattern formation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0048507 | Biological Process | meristem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016020 | Cellular Component | membrane | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 931 aa Download sequence Send to blast |
MMASVEEKIK TGGGMIVGGQ TLAAEMKLLK EMQEHSGVRK TLNSELWHAC AGPLVSLPQV 60 GSLVFYFPQG HSEQVAASTR RTATSQIPNY PNLPYQLLCQ VQNVTLHADK ETDEIYAQMT 120 LQPLNSEREV FPISDFGHKH SKHPSEFFCK TLTASDTSTH GGFSVPRRAA EKLFPPLDYT 180 IQPPTQELVV RDLHDNTWTF RHIYRGQPKR HLLTTGWSLF VGSKRLRAGD SVLFIRDERS 240 QLRVGVRRVN RQQTTLPSSV LSADSMHIGV LAAAAHAAAN RSPFTIFYNP RACPSEFVIP 300 LAKYRKSVFG TQVSVGMRFG MMFETEESGK RRYMGTIVGI SDVDPLRWPG SKWRNIQVEW 360 DEPGCGDKQN RVSVWEIETP ESLFIFPSLT SGLKRPLPSG LLENEWGTLL RRPFIRVPEN 420 GTMELSNSIP NLYSEHMMRM LLKPQLINNN GAFLSAMQQE SAATRGPLQE MKTTLAAENQ 480 MPLKNLHPHS IPDQPNALNM QSLLKNDQPE KLHPLGKIDN HLSSGIVIDK PKSESEVLPD 540 HVIDYPSMEG CNIEKVAANP VNQQGLANQL PFHNQNQSPL LPQSSPWPMP PQIELSMPHP 600 QMIDMVQADS AMVNGLFPQL DINEWMSYAS SQPFAGQNRP TGPLSDLQEH TSLQPQVVNP 660 PLPSMNNEVW DHYVKNLKFL SQADQLTSIC QPGLYGLNGI PSSNNLRDLS AESNNQSEIC 720 VNVDASNSVG TTVVDPSTSS TILDEFCTMK DREFQNPQDC MVGNLSSSQD VQSQITSASL 780 TESHAFPLRD IPDNSGGTSS SHVDFDESSF LQNNSWQQVP APIRTYTKVQ KAGSVGRSID 840 VTTFKNYEEL IRAIECMFGL DGLLNDTKCS GWKLVYVDYE SDVLLVGDDP WEEFVGCVRC 900 IRILSPSEVQ QMSEEGMKLL NSGALQGMNV * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 0.0 | 1 | 382 | 1 | 392 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.41395 | 0.0 | root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: In early embryo and during organ development. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the whole plant with a lower expression in leaves. Detected in embryo axis, provascular tissues, procambium and some differentiated vascular regions of mature organs. {ECO:0000269|PubMed:10476078}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Seems to act as transcriptional activator. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Mediates embryo axis formation and vascular tissues differentiation. Functionally redundant with ARF7. May be necessary to counteract AMP1 activity. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:14973283, ECO:0000269|PubMed:17553903}. | |||||
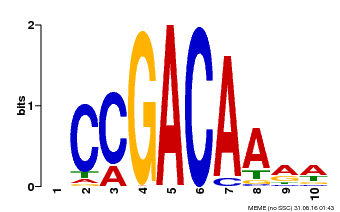
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00153 | DAP | Transfer from AT1G19850 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.14G217700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015036 | 0.0 | AP015036.1 Vigna angularis var. angularis DNA, chromosome 3, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003544394.2 | 0.0 | auxin response factor 5 | ||||
| Swissprot | P93024 | 0.0 | ARFE_ARATH; Auxin response factor 5 | ||||
| TrEMBL | I1MC56 | 0.0 | I1MC56_SOYBN; Auxin response factor | ||||
| STRING | GLYMA14G40540.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8991 | 32 | 41 | Representative plant | OGRP8260 | 12 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19850.1 | 0.0 | ARF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.14G217700.1.p |
| Entrez Gene | 100790289 |



