 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.14G030700.1.p | ||||||||
| Common Name | GLYMA_14G030700, LOC100795553 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 327aa MW: 37654.5 Da PI: 8.7334 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 161.8 | 2.7e-50 | 36 | 163 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
pGfrFhPtdeelv +yL++kve+k+l++ e ik++diyk++PwdLpk + +ekewyfF+ r +ky+++ r+nr+t sg+Wkatg dk+
Glyma.14G030700.1.p 36 LPGFRFHPTDEELVGFYLRRKVEKKPLRI-ELIKQIDIYKYDPWDLPKVSSVGEKEWYFFCIRGRKYRNSIRPNRVTGSGFWKATGIDKP 124
69***************************.89***************8777899************************************ PP
NAM 92 vlsk..kgelvglkktLvfykgrapkgektdWvmheyrl 128
+++ +e +glkk+Lv+y+g+a kg+ktdW+mhe+rl
Glyma.14G030700.1.p 125 IYCVrePQECIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 163
**98545566***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 4.45E-55 | 33 | 194 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 53.373 | 35 | 195 | IPR003441 | NAC domain |
| Pfam | PF02365 | 5.2E-25 | 37 | 163 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 327 aa Download sequence Send to blast |
MKVPKFSSVA HRLIMDVAKL HNSDDDDDKK EEEVVLPGFR FHPTDEELVG FYLRRKVEKK 60 PLRIELIKQI DIYKYDPWDL PKVSSVGEKE WYFFCIRGRK YRNSIRPNRV TGSGFWKATG 120 IDKPIYCVRE PQECIGLKKS LVYYRGSAGK GTKTDWMMHE FRLPPNGKTS NNPQANDAND 180 VQEAEVWTLC RIFKRVPTYK KYTPNLKDSA APITEPNHPT NSSSSKTCSL ESDNCKPYLT 240 FTNSSSPQLI LQNERKPVIN GHVDVERNHL FLGQLGNVAQ APSFWNHHQN NNVEFANENW 300 DDLTSVVQFA IDPSRVSDHS KEFNRF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-48 | 21 | 201 | 3 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-48 | 21 | 201 | 3 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-48 | 21 | 201 | 3 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-48 | 21 | 201 | 3 | 171 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 1e-48 | 21 | 201 | 3 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 1e-48 | 21 | 201 | 3 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, root caps, cotyledons, tips and margin of young leaves, senescent regions of fully expanded leaves and floral tissues, including old sepals, petals, staments, mature anthers and pollen grains. Not detected in the abscission zone of open flowers, emerging lateral roots and root meristematic zones. {ECO:0000269|PubMed:22345491}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
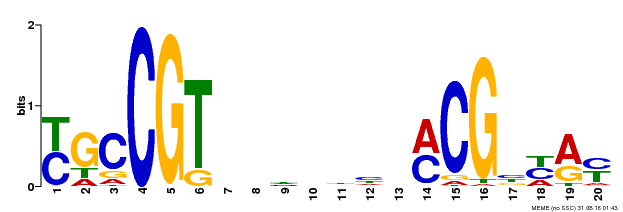
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.14G030700.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT095530 | 0.0 | BT095530.1 Soybean clone JCVI-FLGm-15A17 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006595570.2 | 0.0 | uncharacterized protein LOC100795553 isoform X1 | ||||
| Swissprot | Q9SK55 | 2e-95 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | K7M4P0 | 0.0 | K7M4P0_SOYBN; Uncharacterized protein | ||||
| STRING | GLYMA14G03441.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1013 | 34 | 112 | Representative plant | OGRP17 | 15 | 800 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 7e-96 | NAC domain containing protein 42 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.14G030700.1.p |
| Entrez Gene | 100795553 |



