 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Glyma.03G261800.1.p | ||||||||
| Common Name | GLYMA_03G261800, LCL2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine; Soja
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 749aa MW: 82084.2 Da PI: 6.1436 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.6 | 4.3e-16 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+eE+ ++++a k++G W +I +++g ++t+ q++s+ qk+
Glyma.03G261800.1.p 24 RERWTEEEHNRFLEALKLYGRA-WQRIEEHIG-TKTAVQIRSHAQKF 68
78******************88.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 20.173 | 19 | 73 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-8 | 21 | 71 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.8E-17 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.1E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.1E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.44E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 749 aa Download sequence Send to blast |
MDADSSGEEV VIKTRKPYTI TKQRERWTEE EHNRFLEALK LYGRAWQRIE EHIGTKTAVQ 60 IRSHAQKFFT KLEKEAFVKG VPIGQALDID IPPPRPKRKP SNPYPRKTNV GAPTLHSEAK 120 HGKSLISIAS SHGKQALDLE KEPLPEKHNV DLRPTTVKEN KDGSCLNVFT IIQEAPCSSV 180 SSANKNSTSI SVPLRNSCAL REFIPSVKEV ITRDETNESF VTDELENQKL EIDDGKHTQK 240 TNDTCEVSKL ENSGASELVQ TEKTDGRNCA LTIDGVPGNQ NYPRHVPVHV VDGNLGTSTQ 300 NLSPDMVFQD SIFQPKGGVN RQPNLVTNSA TSHISECQNN AARSSIHQSF PPSPPFAQDD 360 YHSFLHMSST FSSLIVSTLL QNPAAHAAAS FAATFWPYAN AETSADSPVC TPDFPSRQIG 420 SPPSVTAIAA ATVAAATAWW AAHGLLPLCG PLHTAFACPP ASATTVPSMI IDESPQKTER 480 GEIKPQNPPL QDQIPDPEHS EAQHSAPKSP AVSSSKSEDR GDANLDTSPK ATNHEMNQAI 540 SENPDSNKMK GRKPVDRSSC GSNTTSSSEE TELLEKDEKE KEEPKTPDAN VLDTELSNRR 600 SRSISNLTDS WKEVSEEGRL AFQALFSREV LPQSFSPTHH LINKDNQIDS IKDNELNTDY 660 KDEDLESKKC SSICDGVQKN LLFVKDNNEE EGLLTIGLGP GKLKTRRTGF KPYKRCSVEA 720 NENRIGTACI QGEEKGPKRL RLNGEAST* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Gma.17038 | 0.0 | cotyledon| flower| leaf| pod| root | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
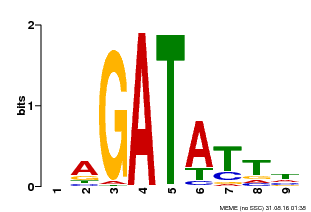
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Glyma.03G261800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU076434 | 0.0 | EU076434.1 Glycine max late elongated hypocotyl and circadian clock associated-1-like protein 2 (LCL2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001236400.1 | 0.0 | MYB transcription factor MYB114 | ||||
| Refseq | XP_025983483.1 | 0.0 | MYB transcription factor MYB114 isoform X1 | ||||
| Refseq | XP_028226910.1 | 0.0 | protein LHY-like | ||||
| Refseq | XP_028226911.1 | 0.0 | protein LHY-like | ||||
| Refseq | XP_028226912.1 | 0.0 | protein LHY-like | ||||
| Refseq | XP_028226913.1 | 0.0 | protein LHY-like | ||||
| TrEMBL | C0SNP2 | 0.0 | C0SNP2_SOYBN; Late elongated hypocotyl and circadian clock associated-1-like protein 2 | ||||
| STRING | GLYMA03G42260.6 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4860 | 25 | 37 | Representative plant | OGRP9903 | 5 | 10 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 2e-81 | circadian clock associated 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Glyma.03G261800.1.p |
| Entrez Gene | 778086 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



