 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D12G1530 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 605aa MW: 65791.8 Da PI: 9.334 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.1 | 3.1e-05 | 89 | 111 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C+ C+k F r nL+ H r H
Gh_D12G1530 89 FICEVCNKGFQREQNLQLHRRGH 111
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.1 | 0.00029 | 165 | 187 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k++ +s+ k H +t+
Gh_D12G1530 165 WKCEKCSKRYAVQSDWKAHSKTC 187
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 6.8E-6 | 88 | 111 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.5E-7 | 88 | 111 | No hit | No description |
| Pfam | PF12171 | 3.0E-5 | 89 | 111 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.0052 | 89 | 111 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.99 | 89 | 111 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 91 | 111 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 130 | 130 | 160 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-5 | 153 | 186 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.5E-7 | 160 | 185 | No hit | No description |
| SMART | SM00355 | 130 | 165 | 185 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 605 aa Download sequence Send to blast |
MATASSSSGP FFGIREENQS QTKQQQQQHT PTSSTGPVDP SPPPKKKRNQ PGTPSKYPIF 60 FFILFLLHIY PDAEVVALSP KTLMATNRFI CEVCNKGFQR EQNLQLHRRG HNLPWKLKQK 120 TTKEVKKKVY LCPEPTCVHH DPSRALGDLT GIKKHYSRKH GEKKWKCEKC SKRYAVQSDW 180 KAHSKTCGTR EYRCDCGTLF SRRDSFITHR AFCDALAQET ARHPPPLNSI GTHLYGTSNY 240 MGLGLSQVGT QISSIQDPNN NQTAADILKL GGGGASRSSQ FDHLLPPSMG SSSSSSFRPQ 300 QQAMGFVMEE SNQIFNQEHQ SQHGLLGNSK PFQGLMQFPP DDDNNNNNNN ASNASSAANL 360 FNLSFLSNSG NNGCAGDYNL TSSGLLMSDN FNNENGGNGG TNNASNLFSN NLMSDQMTSN 420 IPSLFSSSVQ NNNMVPQMSA TALLQKAAQM GSTSSNKNTN DDSLLRTFAN SSSTGTKTSN 480 FGAIFGDTAG NNLHELMNSI ASGNSPIFGS GYDAVHGQES PYTNRSSMGQ EQPPPPPPQS 540 MNVSGGGSDR LTRDFLGVGQ IVRNMRGGAA QQRQGIGISR LSSERNSNIT APTSQQSFGE 600 RGNFQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 1e-34 | 161 | 222 | 3 | 64 | Zinc finger protein JACKDAW |
| 5b3h_F | 1e-34 | 161 | 222 | 3 | 64 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ghi.20456 | 0.0 | boll | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in leaf tissues. {ECO:0000305|PubMed:22898356}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
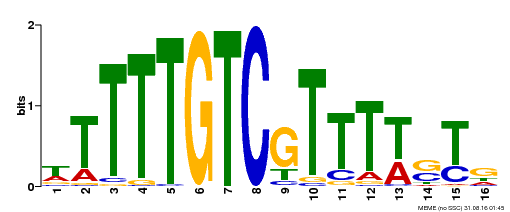
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX614606 | 0.0 | JX614606.1 Gossypium hirsutum clone NBRI_GE58125 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012438385.1 | 0.0 | PREDICTED: protein indeterminate-domain 5, chloroplastic-like | ||||
| Swissprot | Q9ZUL3 | 1e-140 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | A0A0D2T7B8 | 0.0 | A0A0D2T7B8_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.008G169000.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4373 | 26 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02080.1 | 1e-105 | indeterminate(ID)-domain 4 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



