 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_D12G0791 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1088aa MW: 123143 Da PI: 6.4749 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.4 | 4.4e-56 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
l+ ++rwl++ ei++iL n++k+++++e+++ p+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+e+n++fqrr+y
Gh_D12G0791 20 LEAQHRWLRPAEICEILRNYKKFHISSEPAHMPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEQNENFQRRSY 117
5679********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
w+Lee+l++ivlvhy++vk
Gh_D12G0791 118 WMLEEDLSHIVLVHYRDVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.133 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.5E-79 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.5E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.4E-5 | 504 | 591 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 9.96E-19 | 505 | 591 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 2.1E-6 | 505 | 590 | IPR002909 | IPT domain |
| PROSITE profile | PS50297 | 17.237 | 689 | 800 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.03E-12 | 689 | 798 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.0E-15 | 690 | 801 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.64E-15 | 690 | 801 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.992 | 739 | 771 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0062 | 739 | 768 | IPR002110 | Ankyrin repeat |
| Pfam | PF13637 | 4.3E-5 | 741 | 796 | No hit | No description |
| SMART | SM00248 | 1000 | 778 | 807 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.1E-8 | 909 | 964 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.059 | 913 | 935 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.931 | 915 | 943 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.029 | 915 | 934 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0039 | 936 | 958 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.578 | 937 | 961 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 939 | 958 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1088 aa Download sequence Send to blast |
MAESRRYVLT NQLDIDQILL EAQHRWLRPA EICEILRNYK KFHISSEPAH MPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKAGSIDVLH CYYAHGEQNE NFQRRSYWML 120 EEDLSHIVLV HYRDVKGNRT NFNRLKETEG AIPYSQEAVG IVPNSEVESS MSSNFLPNNS 180 QIPSQTMDTA SMNSVHVQAS EYEDAESVSN HQASSQFHSF LDSHQPVAGR ADTRFYDPYV 240 HVSHSNNCYG KPSITAFQLT QTDKDREYNV AGITYEPQKN LDFTSWEDVL GNCDRGVESA 300 QYQPPFTLKQ NDTVGLLFDN SFLKKQAFED QSHAQEEWQG YEGDSSHIVK WSLDQKLHPD 360 LRYDLTSKFD EEVNHNLHPE KQHDHYLLNN QLTDPSKGDH EYVPKPDSEN YLTLEGKSVY 420 SSAMRPHLFD GSLAEEGLKK LDSFNRWMSK ELGDVDESHT HSSSGAYWDE VEGQNGIDVS 480 SIPSQEQLDT FMLGPSLSHD QLFSIIDFSP NWAYVGSEIK VLITGRFLKS QGHAENCKWS 540 CMFGEVEVPA EVIADGVLRC HTPIHEAGRV PFYVTCSNRL ACSEVREFEY RVSHIQDIDT 600 VDNPSNNAIE ILDMRFGRLL CLGSSSPASN TNSIADISQL SNKINSLLKE DVEEWDQMLA 660 HNLEEDFFLE KLKEQLLKKL LKEKLRVWLL QKIVEGGKGP SLLDKGGQGV IHFASALGYD 720 WALEPTIVAG VSVNFRDVNG WTALHWAASS GRERTVASLI SLGAAPGALT DPTPEYPLGR 780 TPADLASANG HKGISGYLAE CDLSSHLLSL NLDKQGSAST TDSRPDVIQK ILELNTAPLN 840 CGDVSDGPSL KDSLAAVRNA TQAAARIHQV FRVQSFQNRQ LKEYGNDKYG MSDERALSLL 900 AVKSNKPGQH DERVHAAASR IQNKFRGWKG RKEFLIIRQR IVKIQAHVRG HQVRKNYRKI 960 VWSVGIVEKV ILRWRRKGSG LRGFKPETLT KGPSVSVPPK EDDYDFLKKG RKQTEERLQK 1020 ALARVKSMAQ NPAGRDQYSR IKNVVTEIQE KVLYDKVLNF AGETTDLDKD LIDLEKLLDE 1080 DTFMHTAP |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Ghi.3380 | 0.0 | boll| ovule| root | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, leaves, carpels, and siliques, but not in stigmas or other parts of the flower. {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
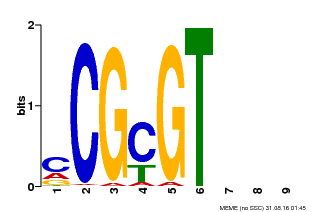
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016738096.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A1U8NGE6 | 0.0 | A0A1U8NGE6_GOSHI; calmodulin-binding transcription activator 3-like isoform X1 | ||||
| STRING | Gorai.008G089900.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8601 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



