 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Gh_A07G2035 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 505aa MW: 56828 Da PI: 5.6933 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.4e-17 | 255 | 301 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g WT++Ed l ++v+++G++ W+ Ia++++ gR +kqc++rw+++l
Gh_A07G2035 255 KGQWTPQEDRVLMQLVTRHGTKKWSQIAKMLN-GRVGKQCRERWHNHL 301
799*****************************.*************97 PP
| |||||||
| 2 | Myb_DNA-binding | 53.3 | 6.5e-17 | 307 | 349 | 1 | 45 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwq 45
+++W++eEd +l+ a+k+ G++ W+ Ia++++ gRt++ +k++w+
Gh_A07G2035 307 KDSWSEEEDMILIAAHKEIGNK-WAEIAKRLP-GRTENTIKNHWN 349
679*******************.*********.***********8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 29.223 | 250 | 305 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.31E-31 | 253 | 348 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.3E-14 | 254 | 303 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-28 | 256 | 308 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 1.60E-13 | 258 | 301 | No hit | No description |
| Pfam | PF13921 | 4.1E-19 | 258 | 315 | No hit | No description |
| PROSITE profile | PS51294 | 19.825 | 306 | 356 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.4E-16 | 306 | 354 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.62E-12 | 309 | 352 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-22 | 309 | 354 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010439 | Biological Process | regulation of glucosinolate biosynthetic process | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1904095 | Biological Process | negative regulation of endosperm development | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 505 aa Download sequence Send to blast |
MEYDTSFKED FPFLSSLFSE NNSNSSFKPD FNGCFPLPDG SSPSSSSSPS SSSSKALVHN 60 FILNQDQDTT PTGNNNNGSF LNNHTHHFHQ FPIDGSSKNP FFEDSTTCTD PFIDPYTNDL 120 NAYIPSLSFT VPDHGTSLNN GLTFQAFSTE SPCWDFSQNK ASAPPEAGDH HRSYQQQQQE 180 PMDFQDQPPT PPPPPPPVAT KVAEDASCIT NQNGRNNNNQ DRDDDDDDDE KNNNNRRFLK 240 AKRVNQATKK NSIIKGQWTP QEDRVLMQLV TRHGTKKWSQ IAKMLNGRVG KQCRERWHNH 300 LRPDIKKDSW SEEEDMILIA AHKEIGNKWA EIAKRLPGRT ENTIKNHWNA TKRRQFTRRS 360 KAKDGNSNPP KGSLLQNYIK SVSSPTELTP AAAASSSSQH ENEFEMTDAN PHMVQLETSS 420 LNSTGWSIGD LNDHVQRQRQ QQQEVNYCFD ANVYNDIHRN QSFGSMLEGD GNGNGSGMGN 480 FELPLEMDSL KKELDLLEMI SQGNL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1mse_C | 7e-45 | 253 | 355 | 2 | 104 | C-Myb DNA-Binding Domain |
| 1msf_C | 7e-45 | 253 | 355 | 2 | 104 | C-Myb DNA-Binding Domain |
| Search in ModeBase | ||||||
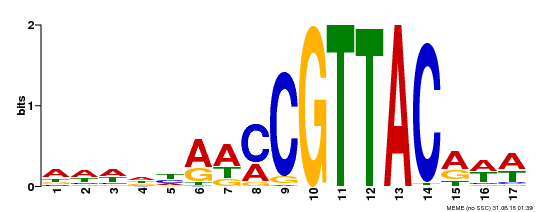
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00385 | DAP | Transfer from AT3G27785 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JX615468 | 0.0 | JX615468.1 Gossypium hirsutum clone NBRI_GE59465 microsatellite sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017644915.1 | 0.0 | PREDICTED: myb-like protein AA | ||||
| TrEMBL | A0A2P5YX08 | 0.0 | A0A2P5YX08_GOSBA; Uncharacterized protein | ||||
| STRING | Gorai.001G263600.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6052 | 26 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G27785.1 | 2e-72 | myb domain protein 118 | ||||



