 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | EPS65995.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Genlisea
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 350aa MW: 40731.2 Da PI: 8.6271 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 21.7 | 5.3e-07 | 64 | 85 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg+sF+ + +Lk+H+ +H
EPS65995.1 64 TCESCGMSFRKPAHLKQHMLSH 85
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 19.9 | 2e-06 | 128 | 152 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ sF+r+++LkrH H
EPS65995.1 128 FICPlkDCNSSFRRNDHLKRHVLQH 152
78*******************9877 PP
| |||||||
| 3 | zf-C2H2 | 16.8 | 1.9e-05 | 157 | 182 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp C+ +F+++ n++rH++ H
EPS65995.1 157 FECPteGCKSRFTTQGNMTRHLKEfH 182
89*********************988 PP
| |||||||
| 4 | zf-C2H2 | 18.7 | 4.7e-06 | 197 | 221 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+kC+ Cgk+F+ +s L++H ++H
EPS65995.1 197 HKCGeeGCGKVFKYSSRLRKHENSH 221
799999****************999 PP
| |||||||
| 5 | zf-C2H2 | 19.3 | 3.2e-06 | 291 | 314 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C+ C+ +Fs+ksnL++Hi+ H
EPS65995.1 291 CSfpECSLTFSTKSNLNQHIKAvH 314
77779***************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 4.24E-6 | 17 | 85 | No hit | No description |
| SMART | SM00355 | 24 | 19 | 42 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 2.0E-5 | 60 | 86 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.367 | 63 | 90 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.031 | 63 | 85 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 65 | 85 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 6.8E-10 | 125 | 154 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.88E-7 | 125 | 154 | No hit | No description |
| SMART | SM00355 | 0.0021 | 128 | 152 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.679 | 128 | 157 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 130 | 152 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-7 | 155 | 182 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 2.31E-6 | 155 | 183 | No hit | No description |
| SMART | SM00355 | 0.0038 | 157 | 182 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.889 | 157 | 187 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 159 | 182 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.6E-7 | 189 | 221 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 8.22E-5 | 196 | 223 | No hit | No description |
| SMART | SM00355 | 0.0093 | 197 | 221 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.554 | 197 | 221 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 199 | 221 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.43 | 229 | 254 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 231 | 254 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 12 | 257 | 278 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.2E-6 | 268 | 311 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 3.14E-9 | 268 | 326 | No hit | No description |
| PROSITE profile | PS50157 | 11.115 | 289 | 319 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0079 | 289 | 314 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 291 | 314 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 6.2 | 320 | 346 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 8.663 | 320 | 350 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 322 | 346 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MGNEGEGRRE AIFKDIRRYY CDYCGISRSK KALITAHVLA QHPEEVKKQQ DCSEGEDCFE 60 KSNTCESCGM SFRKPAHLKQ HMLSHSLMNS KMREAPFLEL DQYQAGDCLG SSIKFSLTTL 120 DKRFERPFIC PLKDCNSSFR RNDHLKRHVL QHEGKTFECP TEGCKSRFTT QGNMTRHLKE 180 FHGEGLPRDE TRKTTEHKCG EEGCGKVFKY SSRLRKHENS HVKLHCVEAM CTEPDCMKYF 240 SNEKCLQAHI QSSHRYITCE KCGSRQLKQN IKRHMRSHES SDPSSPRRIE CSFPECSLTF 300 STKSNLNQHI KAVHLETRRF KCSVPDCDMR FAFKHVRDKH EKSELHTYTS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2i13_A | 5e-13 | 109 | 350 | 4 | 190 | Aart |
| 2i13_B | 5e-13 | 109 | 350 | 4 | 190 | Aart |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
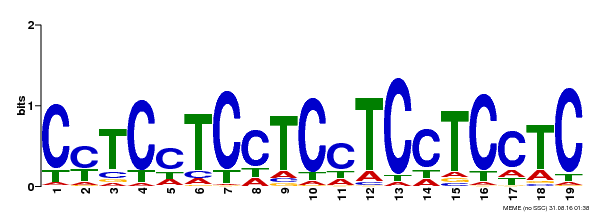
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011079244.1 | 1e-132 | transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 7e-97 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | S8CGP3 | 0.0 | S8CGP3_9LAMI; Uncharacterized protein (Fragment) | ||||
| STRING | VIT_08s0040g00240.t01 | 1e-128 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3644 | 24 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 3e-99 | transcription factor IIIA | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



