 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cotton_A_09671_BGI-A2_v1.0 | ||||||||
| Common Name | F383_05499 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Malvoideae; Gossypium
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1614aa MW: 182626 Da PI: 9.0696 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.8 | 0.00035 | 1515 | 1537 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
Cotton_A_09671_BGI-A2_v1.0 1515 CPvkGCGKKFFSHKYLVQHRRVH 1537
9999*****************99 PP
| |||||||
| 2 | zf-C2H2 | 11.6 | 0.00084 | 1573 | 1599 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
Cotton_A_09671_BGI-A2_v1.0 1573 YVCAeeGCGQTFRFVSDFSRHKRKtgH 1599
89********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.9E-13 | 23 | 64 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.314 | 24 | 65 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 1.7E-13 | 25 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.783 | 199 | 368 | IPR003347 | JmjC domain |
| SMART | SM00558 | 6.9E-49 | 199 | 368 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.46E-25 | 213 | 384 | No hit | No description |
| Pfam | PF02373 | 2.9E-35 | 232 | 351 | IPR003347 | JmjC domain |
| SMART | SM00355 | 26 | 1490 | 1512 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1513 | 1537 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.445 | 1513 | 1542 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 1514 | 1541 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1515 | 1537 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.74E-9 | 1529 | 1571 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-8 | 1542 | 1567 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1543 | 1567 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1543 | 1572 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1545 | 1567 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.45E-8 | 1561 | 1595 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.3E-9 | 1568 | 1596 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.85 | 1573 | 1599 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.783 | 1573 | 1604 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1575 | 1599 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0032259 | Biological Process | methylation | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0008168 | Molecular Function | methyltransferase activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1614 aa Download sequence Send to blast |
MAASSLSPEP SQEVFPWLKY LPFAPEYRPT LVEFQDPIAY IFKIEKEASQ YGICKIIPPV 60 PPASKKAAIG NLNRSLLERA VVNASSDSKP APTFTTRQQQ IGFCPRKQRP VLKSVWQSGE 120 YYTFQEFEAK AKNFERNYLK KYSKKGSLSA LEIETLFWKA TVDQPFDIEY ANDMPGSAFA 180 ALSSKKSSGG GRETGEGVTV AETPWNMRAV SRAKGSLLRF MKEEIPGVTS PMVYIAMLFS 240 WFAWHVEDHD LHSLNYLHMG AGKTWYGVPR DAAVAFEEVV RVDGYAGEFN PLVTFSTLGE 300 KTTVMSPEVF VHAGIPCCRL VQNAGEFVVT FPRAYHSGFS HGFNLGEAAN IATPEWLRVA 360 RDAAIRRASI NYPPMVSHFQ LLYDLALDLC SRVPTNIGAK PKSSRLKDKK KCEGETLVKE 420 LFVQDLIQNN DLLHILGKGS SVVLLPKSST DISLCSESRV ASRLRINPKM SLGLCRYKEA 480 MKPSQVLAFD EIMQCKNEEI KGIKGFYSVK GKTVSTYEVD RDSSFTGADY LCKVPSQTLN 540 ANIERESSVQ GDTLSDQRLF TCVTCGILCF ACIAVLQPTE QAARYLMSAD CSFFNDWTFG 600 SGVTCDRFTA THGDGITSEQ NPSTRGMNKT APNALYDVPV QSVEDKFQMK NQSKEVPEDT 660 KERRNTSALG LLASTYGNSS DSEDDNVEPK ATASGDETNS TNTIPGRKLQ HNDFKEEAPV 720 HVFDCDPEPG SKRSPPTINQ ELVSNGLGDK CSDPTIESHG AEKMRFSKAF TQMENADSPF 780 APNSDEDSSR MHVFCLEHAI EVEQQLRQIG GVHIFLLCHP EYPKIEAEAK LVAEELGIDY 840 LWNNILFGDA TKDDEVRIRS ALDSEDAIPG NGDWAVKLGI NLFYSSNLSH SMLYSKQMPY 900 NSVIYSAFGR NSPTSSPTKL NSYGRRSRKQ KKVVAGKWCG KVWMSNQVHP FLTQRDPGEQ 960 EQEKSYHAWA TSDENLESKP ENIRKAETSK VAKRFTRKSK TRAGTTPSKK AKCIEPESVV 1020 SDDSLDGNSL RQQQRFFRGK KPKLIEKEKE ISYDSLDDDS LLHHRDLSRR KGAKFIEREA 1080 SESEDAEEDS DDQQFRKNLG GKQGKYIVEN DVVSGDSLDK TSAKQYRRIP RSCLAKFMKR 1140 EGSVSADEQE EISHQFHKRI PRGRHIKLFE RRLAVSDDSR ADNSLKQYRR MPKGKRAKFF 1200 EREEAMSDDA SDNDDSQTQH RRIPSGKQMK CMEREDEFSD DSLEDNPQQQ HRRIAQRKVS 1260 KFSDQEDIVS FDSLKGNSHR QHRRIPRSQL TQFIEREDAG SSDSPDDSSL QQLRRILRSK 1320 HAKILEKEDA VSDDSDDTSP QQFRKTPRSR QGKSIESEDV VSYDSSDENY GQPNSSFRSR 1380 KKKAPTPRQT KQKAPKNVKQ GKRRTTKQVI SQQSKQDTRR NRNTKIEQSA RQDNSYDEDE 1440 IEGGPSTRLR KRARKPSKEP ETKPKVKKQA LKNKAKSASN AKVRDEEAEY QCDMEGCTMS 1500 FGLKQELVLH KKNICPVKGC GKKFFSHKYL VQHRRVHMDD RPLKCPWKGC KMTFKWAWAR 1560 TEHIRVHTGA RPYVCAEEGC GQTFRFVSDF SRHKRKTGHS LKKGKEIKHE SRKL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 5e-73 | 17 | 389 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 5e-73 | 17 | 389 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
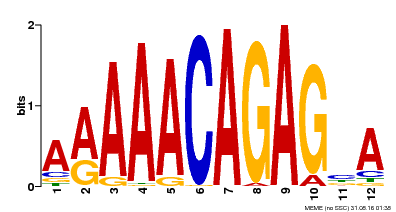
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017611137.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A0B0P356 | 0.0 | A0A0B0P356_GOSAR; Lysine-specific demethylase REF6-like protein | ||||
| STRING | Gorai.006G270100.1 | 0.0 | (Gossypium raimondii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



