 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | mrna09903.1-v1.0-hybrid | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1493aa MW: 165633 Da PI: 9.4815 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11 | 0.0013 | 1373 | 1398 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++C C++sF +k++L+ H ++ +
mrna09903.1-v1.0-hybrid 1373 FVCDieGCTMSFGTKHELNLHKKNvC 1398
899999***************99866 PP
| |||||||
| 2 | zf-C2H2 | 13.5 | 0.00022 | 1397 | 1420 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp Cgk F ++ +L++H r+H
mrna09903.1-v1.0-hybrid 1397 VCPvkGCGKKFFSHKYLVQHRRVH 1420
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.8 | 0.00073 | 1456 | 1482 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
mrna09903.1-v1.0-hybrid 1456 YVCAepGCGQTFRFVSDFSRHKRKtgH 1482
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.8E-16 | 20 | 61 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.243 | 21 | 62 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 3.7E-15 | 22 | 55 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 1.54E-24 | 115 | 174 | No hit | No description |
| PROSITE profile | PS51184 | 32.547 | 187 | 366 | IPR003347 | JmjC domain |
| SMART | SM00558 | 5.8E-46 | 197 | 366 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 1.54E-24 | 215 | 365 | No hit | No description |
| Pfam | PF02373 | 3.5E-36 | 230 | 349 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-4 | 1369 | 1394 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 14 | 1373 | 1395 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.612 | 1396 | 1425 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0037 | 1396 | 1420 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.33E-5 | 1396 | 1432 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.7E-6 | 1396 | 1419 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1398 | 1420 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.2E-9 | 1420 | 1448 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.637 | 1426 | 1455 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0017 | 1426 | 1450 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1428 | 1450 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.27E-9 | 1436 | 1480 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.1E-9 | 1449 | 1479 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.62 | 1456 | 1482 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.803 | 1456 | 1487 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1458 | 1482 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1493 aa Download sequence Send to blast |
MAAPPEQPPE VLPWLRALPV APEYHPTWAE FQDPIAYIFK IEKEASQYGI CKIVPPVPPA 60 PKKTAIANLN KSLILRNGPV TGKGPKAQPT FTTRQQQIGF CPRKARPVQR PVWQSGEHYT 120 FSQFEAKAKS FEKSYLKKQR KKGGLSALDI ETLYWKATVD KPFSVEYAND MPGSAFVPLS 180 SKKSGGSTSR EAGDGVTLGE TAWNMRGVSR SRGSLLRFMK EEIPGVTCPM VYVAMMFSWF 240 AWHVEDHDLH SLNYLHMGAG KTWYGVPREA AVAFEEVVRV QGYGGEINPL VTFATLGEKT 300 TVMSPEVFIS SGIPCCRLVQ NAGEFVVTFP RAYHTGFSHG FNCGEAANIA TPEWLRVAND 360 AAVRRASINY PPMVSHFQLL YDLALALCSR TPVHSSAEPR SSRLKDKKKG EGETVVKGLF 420 VKNVIQNNEL LHVLGKGSSI VLLPQSSSDI SVCSKLRVGS QLRVNPDDLI IDGNRGIKQV 480 SVKGKLASLC ESSRHLSLNG NDSAATPSKM LNMSAKRESN VEGEGLSDQR LFSCVTCGIL 540 SFSCVAIIQP REAAARYLMS ADCSFFNDWA VDCEPIQGAN GDPNSSKKGP CTETGLKQKS 600 APDSLYDAPF QSADNQNQIT DPSNEVDSNT ENQRDTNALG LLALTYGVSS DSEEDQANQD 660 VPVCGDKSNL SDCSLEGRYE YQSASPPLRA SYGGTAGVRS PTSPGFDCGI GLPTIDGNGL 720 PTIDVYVENR PEATNFKDKG HQYSVDLDTN NLALTKTNGL VGTSIDPMKV SYSGSPDAFD 780 VQPTGFGQVT LRKDSTGTSF APGFDHDSSR MHVFCLEHAV EVEQQLRSFG GAHILLLCHP 840 DYPRIVDEAK EIAEELGVNY PWNDLVFRNA TRADEQRIQS ALDSEEAIAG NGDWAVKMGI 900 NLFYSASLSR SHLYSKQMPY NSVIYNAFGR SSPATSPAGP EVCGRRPAKQ KKVVVGKWCG 960 KVWMSNQVHP FLIKREHEEK KVEQERRRFQ ESPIPDEKLH GNTESTHKTE KTVVTKQYSR 1020 KRKMTVDGET TKKAKRTDAV SAQSVDDDSH LQQMRFLKNK QGKHIESGPT KKSKIEKEDA 1080 VSSDSMEDDF RQQNRRTLRS KQAKHSVGDD DVSDDSMGVD SQQQQTRIAK SKQAKHSAKD 1140 FSVVSDDSVG VDSDHQQKRV AESNTREFSA VSDDSLDESI HQLHRRSLRR NKGKSIGREN 1200 FTSQNLYGVS SRQKQKKTSK SKQAKIVERE EAALDETTDD NAALQHKIVR GKQIKPETLQ 1260 QMKRETPHRV RQGSRRLQES QQQTPRIRNT TDVHAEEPEG GPSTRLRKRP PKEQPETSRK 1320 KAKVQPETGR KKAKEQQQTG RIKVNTASAV KTKNASARKT KNASGARVEE AEFVCDIEGC 1380 TMSFGTKHEL NLHKKNVCPV KGCGKKFFSH KYLVQHRRVH EDDRPLRCPW KGCKMTFKWA 1440 WARTEHIRVH TGARPYVCAE PGCGQTFRFV SDFSRHKRKT GHSVKKGKGR SR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-73 | 12 | 387 | 6 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-73 | 12 | 387 | 6 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1018 | 1023 | SRKRKM |
| 2 | 1318 | 1331 | RKKAKVQPETGRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
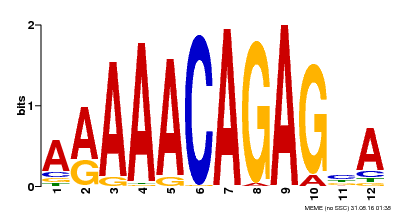
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | mrna09903.1-v1.0-hybrid |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004301036.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A2P6P5A5 | 0.0 | A0A2P6P5A5_ROSCH; Putative [histone H3]-lysine-36 demethylase | ||||
| STRING | XP_004301036.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | mrna09903.1-v1.0-hybrid |



