 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | FANhyb_rscf00000070.1.g00022.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Rosoideae; Potentilleae; Fragariinae; Fragaria
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1981aa MW: 222116 Da PI: 6.4392 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 103.7 | 1.3e-32 | 67 | 141 | 2 | 76 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvg 76
l+ k+rwl++ ei++iL+n++k+++++e++++p+ gsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK
FANhyb_rscf00000070.1.g00022.1 67 LEAKHRWLRPAEICEILQNYKKFHISTEPASTPPGGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKKT 141
6679********************************************************************843 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 47.238 | 62 | 274 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 6.0E-42 | 65 | 269 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 2.4E-27 | 68 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM00429 | 0.52 | 620 | 708 | IPR002909 | IPT domain |
| Gene3D | G3DSA:2.60.40.10 | 1.3E-5 | 622 | 708 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 2.33E-18 | 623 | 708 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 1.4E-6 | 623 | 707 | IPR002909 | IPT domain |
| PROSITE profile | PS50297 | 18.139 | 788 | 927 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.51E-12 | 803 | 915 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.4E-31 | 804 | 918 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.02E-16 | 806 | 917 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.217 | 856 | 888 | IPR002110 | Ankyrin repeat |
| Pfam | PF00023 | 0.012 | 856 | 887 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.032 | 856 | 885 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 260 | 895 | 924 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.45E-7 | 1016 | 1074 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 23 | 1023 | 1045 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.089 | 1025 | 1053 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0017 | 1046 | 1068 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.597 | 1047 | 1071 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 1049 | 1068 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00429 | 1.1 | 1344 | 1432 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.96E-16 | 1346 | 1432 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 3.8E-4 | 1347 | 1431 | IPR002909 | IPT domain |
| Gene3D | G3DSA:2.60.40.10 | 9.9E-4 | 1347 | 1432 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00204 | 1.26E-11 | 1533 | 1645 | No hit | No description |
| PROSITE profile | PS50297 | 15.565 | 1534 | 1657 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 3.4E-31 | 1543 | 1649 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.35E-15 | 1544 | 1645 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 3.5E-6 | 1558 | 1648 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.038 | 1586 | 1615 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.816 | 1586 | 1618 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 480 | 1625 | 1654 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.22 | 1777 | 1799 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.535 | 1778 | 1802 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.035 | 1780 | 1798 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1981 aa Download sequence Send to blast |
MAETKRYGLG NQLGQFLLID FCIKFSSLRI FVILALVLRS ICSFLLAYLL STCRAPAVQL 60 DIQQILLEAK HRWLRPAEIC EILQNYKKFH ISTEPASTPP GGSLFLFDRK VLRYFRKDGH 120 NWRKKKDGKT VKEAHERLKK TDHFQVLMEI WVNLRLRLVC RLIKANELSS LSFLLASAEL 180 QLNLETDIRL RDDADRNFCL ENRGEESCLN VHEVGGLEVL MCCTVTMPTE KTTKIFKDAV 240 IGCLKILNPE GCLLMDLSHI VLVHYREVKG NRTNFNHVKE TEGVAYSNGA EQSARQSEME 300 NSVSSSFNPS SYQMHSQTTE ATSLSSAQAS EFEDAESAFY NQASSRLQPM AEKINSEFAD 360 AYYPTFSSKF DSFNSLSQAY KGEDSIHAGI TYEPRKDRDF ALWDDMENSA TGVQSFQPSF 420 SATHSDTMGS FPKQEIETIG HLYTDSFDKR LVYGMENRPK VQQSWQTSEG SSNWPMDQSI 480 QSHAQYNVTS KLHDGADATD LLKSWPFLMD SDKQNDLQFH LSNTDSISKR NDIIEGKADY 540 PSAIKPLLDG AFGDGLKKLD SFNRWMSKEL EDVDEPQMQS SSGAYWETVE SENEVDESSV 600 PLQVRLDSYM LGPSLSHDQL FSIVDFSPSW AYENSEIKVL ITGRFLKSQH AESCKWSCMF 660 GEVEVPAEVI ADGVLRCYTP IHKAGRVPFY VTCSNRLACS EVREFEYRVA ETQDVDCKDY 720 YSDFSNETLS MRFGNFLTLS STSPNCDPAS IAENSEVNSK ITSLLKNDND EWDKMLQLTS 780 DEDFSLKRVE EQLHQQLLKE KLHAWLLQKL AAGGKGPNVL DEGGQGVLHF GAALGYDWVL 840 LPTITAGVSV NFRDVNGWTA LHWAAFCGRE RTVASLISLG AAPGALTDPT AKYPSGETPA 900 DLASEQGHKG IAGYLAESAL SKHLESLNLD LKDGNSAEIS GAKAVSGSSR DGELTDGLSL 960 RDSLTAVCNA TQAAARIHQV FRVQSFQRKQ LKEYGGDKFG ISNERALSLI AVKSHKSGKR 1020 DEHVDAAAVR IQNKFRSWKG RKDFLIIRQR IVKIQAHVRG HQVRKNYKKI VWTVGIVEKI 1080 ILRWRRKGSG LRGFKPEPLT EGPSMQVSST KEDDDDVLKE GRKQTEERMQ KALARVKSMA 1140 QYPEARDQYR RLLNVVTEIQ ETKVLNSSEG TSAYMDDDLI DIEALFDDDV FMPTATSRLS 1200 HIVLVHYRDV KTSEGNLSCS SELSINRNVH SDTSYHASTS FPRGVNADEL LNSLGPPCHM 1260 DLIKQNESSL HRHSDNKNDP WRLKNLDSAN QCMNTDLGDL TEPHMSSSED HSNTIASENE 1320 IDGFSALKQV EIATLGSSLF QDLLFNIMDF SPNWAYERAE TKVVVVGRFL KSQELDSFNW 1380 SCMFGEVEVP AEVVADGVLR CYTPIHRAAR IPFYVTCSNR LACSEVKIFE YRVNHIQDAD 1440 FKDNCCEENT LVMRFGKLLS PSVMYPSFNP RVKHRQYDPI CVSDNSDLTR KISSLLKNGD 1500 EERAEMLKLF PDAFSLDTVK EQLLQQFLKD KFHEWLLKKL AVGGNGLRVL DEGGQGVLHI 1560 GAALGYDWVL SPMKIAGVSI NFRDVNGWTA LHWAAFCGRD RAVALLISLG GAPGALTDPC 1620 TKNPTGKTPA DLASENGYKG IAGYLAASSL SAHISSLKLD IKDAHSAESS REKSVQPDGM 1680 STHCSNGDLA DELSMRVCST AVCDPIQASA RIHQVFSVQP FQRKQLKEYG DKSKTSDEGA 1740 LSLIAVKSHK KCDEHVSCAA IQTPDKFRNR NIKDFLIIRS RILKIQANVR GHQERKSYTK 1800 ILWAVGILEK IILRWRRKRS GLRGFKREPP SDEGRSIQVS SSRENDYDFL KEGRKQTEER 1860 LQNALARVKS MVQIPEARDQ YNQLSNIFTE IQETKVVDSS EGESTDMDYN GMLQVVNNSI 1920 GISADMDYNI IDTNALLDDN ILRIRTIIIL KPFTGKVKVA KSTTINLHDK LTSLAVDGSK 1980 S |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 3e-14 | 623 | 708 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_B | 3e-14 | 623 | 708 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_C | 3e-14 | 623 | 708 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_D | 3e-14 | 623 | 708 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_E | 3e-14 | 623 | 708 | 9 | 89 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
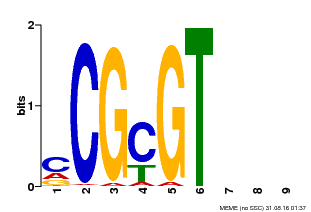
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004288193.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3-like | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A2P6RQI8 | 0.0 | A0A2P6RQI8_ROSCH; Putative transcription factor CG1-CAMTA family | ||||
| STRING | XP_004288193.1 | 0.0 | (Fragaria vesca) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



