 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462956324 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 291aa MW: 31917.9 Da PI: 5.4173 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.2 | 6.8e-19 | 67 | 120 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++ ++q+++Le+ Fe +++++ e+++++A+ l+L rqV vWFqNrRa++k
462956324 67 KKRRLAADQVRALERSFEADNKLDPERKARIARDLRLHPRQVAVWFQNRRARWK 120
4557899**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 126.9 | 8.7e-41 | 66 | 158 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl+++qv++LE+sFe+++kL+perK+++ar+L+l+prqvavWFqnrRAR+ktkq+E+dy aL+++ydal++++++L++++++L +e++e
462956324 66 EKKRRLAADQVRALERSFEADNKLDPERKARIARDLRLHPRQVAVWFQNRRARWKTKQMERDYVALRHSYDALRADHDALRRDKDALLAEIRE 158
69*************************************************************************************999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-19 | 45 | 123 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.56E-18 | 61 | 124 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.422 | 62 | 122 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.1E-18 | 65 | 126 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.48E-16 | 67 | 123 | No hit | No description |
| Pfam | PF00046 | 3.4E-16 | 67 | 120 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 97 | 120 | IPR017970 | Homeobox, conserved site |
| Pfam | PF02183 | 2.9E-16 | 122 | 162 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 291 aa Download sequence Send to blast |
MKRPASRGSS VPVISQETPE QQEDSSVRYS MMEREDVVAG AGAYNEEEEE DEEDDDLGGG 60 RGGLGEKKRR LAADQVRALE RSFEADNKLD PERKARIARD LRLHPRQVAV WFQNRRARWK 120 TKQMERDYVA LRHSYDALRA DHDALRRDKD ALLAEIRELR EKAEKQMAVK PRPLAATTAA 180 AAAVYKQDGS TDSDSSAVFN EEASPYSGGA AATATAFDHH HNPHLHPSFA GFTSFLASSG 240 ALSSSFPSMY SGGGSHLDQE GDHGLLGAAD GFFAEDHHGT GLGSWYGGEG W |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 114 | 122 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
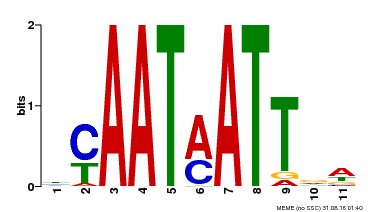
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00587 | DAP | Transfer from AT5G65310 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462956324 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FP099753 | 3e-98 | FP099753.1 Phyllostachys edulis cDNA clone: bphyem212a11, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025792352.1 | 1e-102 | homeobox-leucine zipper protein HOX8-like isoform X1 | ||||
| Swissprot | Q338Z7 | 7e-86 | HOX8_ORYSJ; Homeobox-leucine zipper protein HOX8 | ||||
| TrEMBL | A0A0A9JEK4 | 1e-104 | A0A0A9JEK4_ARUDO; Uncharacterized protein | ||||
| STRING | Pavir.J35191.1.p | 1e-107 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4283 | 31 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G65310.2 | 7e-37 | homeobox protein 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



