 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G65310.2 | ||||||||
| Common Name | ATHB5, ATHB-5, HB5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 294aa MW: 32846.5 Da PI: 4.5293 | ||||||||
| Description | homeobox protein 5 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.4 | 1.2e-18 | 56 | 107 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV +WFqNrRa++k
AT5G65310.2 56 RRLGVEQVKALEKNFEIDNKLEPERKVKLAQELGLQPRQVAIWFQNRRARWK 107
467789*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.4 | 1.5e-41 | 53 | 145 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl eqvk+LE++Fe ++kLeperKv+la+eLglqprqva+WFqnrRAR+ktkqlE+dy +Lk+++dalk+++++L++++++L ++ke
AT5G65310.2 53 EKKRRLGVEQVKALEKNFEIDNKLEPERKVKLAQELGLQPRQVAIWFQNRRARWKTKQLERDYGVLKSNFDALKRNRDSLQRDNDSLLGQIKE 145
69*************************************************************************************999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.84E-19 | 45 | 111 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.41 | 49 | 109 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 5.3E-18 | 52 | 113 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 5.17E-17 | 54 | 110 | No hit | No description |
| Pfam | PF00046 | 5.7E-16 | 55 | 107 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 8.6E-20 | 58 | 116 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 7.0E-6 | 80 | 89 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 84 | 107 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 7.0E-6 | 89 | 105 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 5.1E-17 | 109 | 150 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009787 | Biological Process | regulation of abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000003 | anatomy | whole plant | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0003000 | anatomy | transition zone | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009001 | anatomy | fruit | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007057 | developmental stage | seed germination stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MILTDKQISP RPTTTGFLYS GAGDYSQMFD ALEDDGSLED LGGVGHASST AAEKKRRLGV 60 EQVKALEKNF EIDNKLEPER KVKLAQELGL QPRQVAIWFQ NRRARWKTKQ LERDYGVLKS 120 NFDALKRNRD SLQRDNDSLL GQIKELKAKL NVEGVKGIEE NGALKAVEAN QSVMANNEVL 180 ELSHRSPSPP PHIPTDAPTS ELAFEMFSIF PRTENFRDDP ADSSDSSAVL NEEYSPNTVE 240 AAGAVAATTV EMSTMGCFSQ FVKMEEHEDL FSGEEACKLF ADNEQWYCSD QWNS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 101 | 109 | RRARWKTKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.19698 | 0.0 | flower| inflorescence| leaf| root| seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 247191_at | 0.0 | ||||
| Expression Atlas | AT5G65310 | - | ||||
| AtGenExpress | AT5G65310 | - | ||||
| ATTED-II | AT5G65310 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Localized primarily to the hypocotyl of germinating seedlings. {ECO:0000269|PubMed:12678559}. | |||||
| Uniprot | TISSUE SPECIFICITY: Widely expressed. {ECO:0000269|PubMed:16055682}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a class I HDZip (homeodomain-leucine zipper) protein that is a positive regulator of ABA-responsiveness, mediating the inhibitory effect of ABA on growth during seedling establishment. | |||||
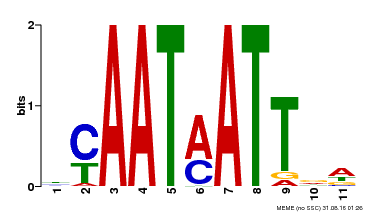
| UniProt | Probable transcription factor that acts as a positive regulator of ABA-responsiveness, mediating the inhibitory effect of ABA on growth during seedling establishment. Binds to the DNA sequence 5'-CAATNATTG-3'. {ECO:0000269|PubMed:12678559}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00587 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G65310.2 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by abscisic acid (ABA) and by salt stress. {ECO:0000269|PubMed:12678559, ECO:0000269|PubMed:16055682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | abscisic acid | |||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT5G65310 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G65310 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF325054 | 0.0 | AF325054.2 Arabidopsis thaliana AT5g65310 (AT5g65310/MNA5_4) mRNA, complete cds. | |||
| GenBank | AY074293 | 0.0 | AY074293.1 Arabidopsis thaliana putative homeobox-leucine zipper protein ATHB-5 (At5g65310) mRNA, complete cds. | |||
| GenBank | AY091340 | 0.0 | AY091340.1 Arabidopsis thaliana putative homeobox-leucine zipper protein ATHB-5 (HD-zip protein ATHB-5) (At5g65310) mRNA, complete cds. | |||
| GenBank | X67033 | 0.0 | X67033.1 A.thaliana mRNA Athb-5. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001032151.1 | 0.0 | homeobox protein 5 | ||||
| Swissprot | P46667 | 0.0 | ATHB5_ARATH; Homeobox-leucine zipper protein ATHB-5 | ||||
| TrEMBL | F4KHX1 | 0.0 | F4KHX1_ARATH; Homeobox protein 5 | ||||
| STRING | AT5G65310.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G65310.2 |
| Entrez Gene | 836656 |
| iHOP | AT5G65310 |
| wikigenes | AT5G65310 |



