 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462878696 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 587aa MW: 66322.5 Da PI: 4.8537 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 110.5 | 1.2e-34 | 178 | 269 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
Fl k+ye++ed+++++++sw g sfvv+++ +f++++LpkyFkh+nf+SF+RQLn+YgF+k++ e+ weF+++ F +g+++ll++i+
462878696 178 FLIKTYEMVEDPATNHVVSWGPGGASFVVWNPPDFSRDLLPKYFKHNNFSSFIRQLNTYGFRKIDPER---------WEFANEDFIRGHTHLLKNIH 265
9****************************************************************999.........******************** PP
XXXX CS
HSF_DNA-bind 99 rkks 102
r+k
462878696 266 RRKP 269
*985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46785 | 4.22E-36 | 173 | 267 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 9.9E-63 | 174 | 267 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene3D | G3DSA:1.10.10.10 | 1.1E-38 | 174 | 267 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 1.0E-30 | 178 | 267 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-19 | 178 | 201 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-19 | 216 | 228 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 217 | 241 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 1.6E-19 | 229 | 241 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 587 aa Download sequence Send to blast |
LLTRSSSRVS LACLPPAATL VELPACVTVV VSVRQESQSS HDPIVCEKKW PDRIGANSRD 60 LVRFWEVIFL HQVVLFSPLL LLVQVELDRP KGRCLYGTSV RISATVECEL GATSPFWWWL 120 RLWEAALRPS YSLHPSLVGL VDTDRPLALL GRETEDAGME ASQSHGGGGG GSSSPPPFLI 180 KTYEMVEDPA TNHVVSWGPG GASFVVWNPP DFSRDLLPKY FKHNNFSSFI RQLNTYGFRK 240 IDPERWEFAN EDFIRGHTHL LKNIHRRKPV HSHSLQNQVN GPLAESERRE YEDEINRLKY 300 EKSLLLADLQ RQNQQHSGIN WQMQLLEDRL MQMEQRQKNI IASLSDILQR NGVASGTLLE 360 TDHFSKKRRV PKIDFCVNDP TIEEQQVPFL QVMGAVTETP SMFPEPFDRM EMSLISLEKF 420 IQRASDESGE DMFTGSAEPS PGVTLGQMHP EPVETNINLQ ESSCYVQSPV LPLPDLEEDA 480 HRTAEADMNS DTTTADTSQD EATTETGLSH EPTKVNDIFW ERFLTETPKF YRADEAESGR 540 QGEECKTEPI EAKEDLKIAV DCSFLHHRDK VDQITQQMGQ LASAENA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 7e-25 | 166 | 267 | 17 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 7e-25 | 166 | 267 | 17 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 7e-25 | 166 | 267 | 17 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
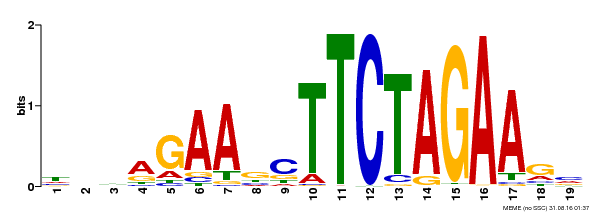
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462878696 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat stress. {ECO:0000269|PubMed:12032317}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004961415.1 | 0.0 | heat stress transcription factor A-4d | ||||
| Swissprot | Q93VB5 | 1e-168 | HFA4D_ORYSJ; Heat stress transcription factor A-4d | ||||
| TrEMBL | K3Z657 | 0.0 | K3Z657_SETIT; Uncharacterized protein | ||||
| STRING | Si022026m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3296 | 37 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 6e-74 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



