 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10011493m | ||||||||
| Common Name | EUTSA_v10011493mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 437aa MW: 47745.3 Da PI: 9.195 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 50.2 | 5.6e-16 | 349 | 392 | 5 | 48 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkke 48
+r+rr++kNRe+A rsR+RK+a++ eLe ++++L++eN++L+ +
Thhalv10011493m 349 RRQRRMIKNRESAARSRARKQAYTVELEAEIAKLKEENDELQRK 392
79*************************************99854 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 7.8E-16 | 342 | 392 | No hit | No description |
| SMART | SM00338 | 1.5E-15 | 345 | 409 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.978 | 347 | 392 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 6.63E-26 | 349 | 394 | No hit | No description |
| SuperFamily | SSF57959 | 3.19E-12 | 349 | 403 | No hit | No description |
| Pfam | PF00170 | 3.1E-13 | 349 | 392 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 352 | 367 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 437 aa Download sequence Send to blast |
MVRRQRDTWL RRQKNELETF LGNGLSQKTK ILQNIIFKNN SMGNEPPGDG GGGGALTRPA 60 LTRQGSIYSL TFDEFQSSLG KDFGSMNMDE LLKNIWSAEE TQAMAMAAST SSMIPVPGGQ 120 EGLQLQRQGS LTLPRTLSTK TVDQVWKDLS KDWNSVGGTS LSQSQSQNQS QSQSQSQSQR 180 QQTLGEVTLE EFLVRAGVVR EEAQVAAKDK DGYFGNDANA GFSVQASPRV VPGLMENLGV 240 ETANHLQVQG SSLPLNVNGA RSTYQQQPIL PKQPGFGYGT QIAQLNSPVV RGGLMGLGDQ 300 PLTNNMGFVQ DGIGKNNGDS SSLSPSPYMF NGGVRGRKSG GTVEKVVERR QRRMIKNRES 360 AARSRARKQA YTVELEAEIA KLKEENDELQ RKQVIANHIC SYLTFISFSF SPYSLCFFTG 420 KDYRNAKESG DGDEES* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in ABA and stress responses and acts as a positive component of glucose signal transduction. Functions as transcriptional activator in the ABA-inducible expression of rd29B. Binds specifically to the ABA-responsive element (ABRE) of the rd29B gene promoter. {ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
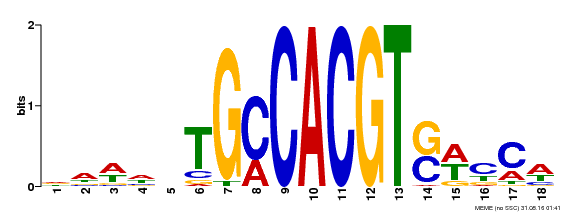
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10011493m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA), cold and glucose. {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:11005831, ECO:0000269|PubMed:15361142, ECO:0000269|PubMed:16284313, ECO:0000269|PubMed:16463099}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | JQ971972 | 0.0 | JQ971972.1 Eutrema salsugineum abscisic acid responsive elements-binding factor 2 (ABF2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024006260.1 | 0.0 | ABSCISIC ACID-INSENSITIVE 5-like protein 5 isoform X1 | ||||
| Swissprot | Q9M7Q4 | 1e-168 | AI5L5_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| TrEMBL | V4KLX3 | 0.0 | V4KLX3_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006393641.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM734 | 27 | 129 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 1e-137 | abscisic acid responsive elements-binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10011493m |
| Entrez Gene | 18010955 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



