 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10001240m | ||||||||
| Common Name | EUTSA_v10001240mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 324aa MW: 36759.3 Da PI: 8.0663 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 104.2 | 7e-33 | 179 | 236 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+Dgy+WrKYGqK vk+s++prsYYrCt+++C+vkk+vers +dp+vv++tYe++Hnh+
Thhalv10001240m 179 EDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSYQDPTVVITTYESQHNHP 236
8********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF038130 | 2.4E-100 | 1 | 323 | IPR017396 | PWRKY transcription factor, group IIc |
| Gene3D | G3DSA:2.20.25.80 | 2.3E-34 | 163 | 238 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 4.71E-29 | 170 | 238 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 32.282 | 173 | 238 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.1E-38 | 178 | 237 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.0E-25 | 179 | 236 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0051607 | Biological Process | defense response to virus | ||||
| GO:0070301 | Biological Process | cellular response to hydrogen peroxide | ||||
| GO:1901002 | Biological Process | positive regulation of response to salt stress | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 324 aa Download sequence Send to blast |
MSNETKDLKN YHYTSSYNHY NTNQNIVNLP YVSGPSTYNA NMVSSQISSD LHSSPQGAYG 60 SGFELSPTLN EFLYSSIDQE NGFYNAYTYN NCQKGDEVVG GGGAIIKSES RVSASPSSSE 120 ADHHHVEDSG KSLRKREAAD GGEADQRSQK VVKTKKKEEK KQREPRVSFM TKTEIDHLED 180 GYRWRKYGQK AVKNSPYPRS YYRCTTQKCN VKKRVERSYQ DPTVVITTYE SQHNHPIPTS 240 RRMGTFSGPA ASEYNSSSLS PVSDFIINNT PRSFSHDDLF RAPYASVNVN PNYQQQQNQD 300 ILRESEFEFL KEMFPSVFFK QEP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 4e-28 | 168 | 238 | 7 | 77 | Probable WRKY transcription factor 4 |
| 2lex_A | 4e-28 | 168 | 238 | 7 | 77 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-TTGAC[CT]-3'), a frequently occurring stress-responsive cis-acting element. Functions as positive regulator of salt stress response. Binds the W box of LTI78/RD29A stress-response gene and directly regulates its transcription under salt stress. Functions antagonistically with VQ9 to regulate sodium and potassium homeostasis under salt stress by regulating the expression of downstream SOS (SALT OVERLY SENSITIVE) stress-responsive genes. The DNA-binding activity of WRKY8 is decreased by VQ9 (PubMed:23451802). Functions as negative regulator of basal resistance to the bacterial pathogen P. syringae and as positive regulator of resistance to the fungal pathogen to B. cinerea (PubMed:20367464). Functions as positive regulator of defense response againt tobamovirus (TMV) by regulating both the abscisic acid and ethylene signaling pathways. Positively regulates ABI4 expression and negatively modulates ACS6 and ERF104 expression by directly binding to the W box consensus motifs within their promoters (PubMed:23650359). {ECO:0000269|PubMed:20367464, ECO:0000269|PubMed:23451802, ECO:0000269|PubMed:23650359}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00545 | DAP | Transfer from AT5G46350 | Download |
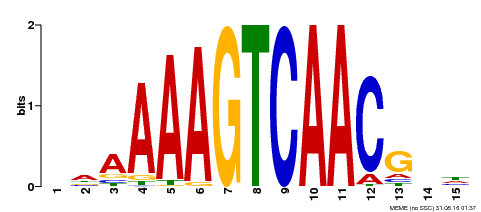
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10001240m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By wounding, abscisic acid (ABA), salicylic acid (SA), H(2)O(2), and infection with P.syringae pv. tomato DC3000 and B.cinerea (PubMed:20367464). Induced by salt stress (PubMed:23451802). {ECO:0000269|PubMed:20367464, ECO:0000269|PubMed:23451802}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006398330.1 | 0.0 | probable WRKY transcription factor 8 | ||||
| Swissprot | Q9FL26 | 1e-163 | WRKY8_ARATH; WRKY transcription factor 8 | ||||
| TrEMBL | V4LJ10 | 0.0 | V4LJ10_EUTSA; WRKY transcription factor | ||||
| STRING | XP_006398330.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1524 | 26 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46350.1 | 1e-129 | WRKY DNA-binding protein 8 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10001240m |
| Entrez Gene | 18015993 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



